This article is meant to be read in a browser, but the web page can take several minutes to load because MathJax is slow. The PDF version is quicker to load, but the latex generated by Pandoc is not as beautifully formatted as it would be if it were from bespoke \({\small{\LaTeX}}\).
This is my attempt to teach myself the backpropagation algorithm for neural networks. I don’t try to explain the significance of backpropagation, just what it is and how and why it works.
There is absolutely nothing new here. Everything has been extracted from publicly available sources, especially Michael Nielsen’s free book Neural Networks and Deep Learning – indeed, what follows can be viewed as documenting my struggle to fully understand Chapter 2 of this book.
I hope my formal writing style – burnt into me by a long career as an academic – doesn’t make the material below appear to be in any way authoritative. I’ve no background in machine learning, so there are bound to be errors ranging from typos to total misunderstandings. If you happen to be reading this and spot any, then feel free to let me know!
The sources listed below are those that I noted down as being particularly helpful.
Many thanks to the authors of these excellent resources, numerous Wikipedia pages and other online stuff that has helped me. Apologies to those whom I have failed to explicitly acknowledge.
A neural network is a structure that can be used to compute a function. It consists of computing units, called neurons, connected together. Neurons and their connections contain adjustable parameters that determine which function is computed by the network. The values of these are determined using machine learning methods that compare the function computed by the whole network with a given function, \(g\) say, represented by argument-result pairs – e.g. \(\{g(I_1)=D_1,g(I_2)=D_2,\ldots\}\) where \(I_1\), \(I_2\),\(\ldots\) are images and \(D_1\), \(D_2\),\(\ldots\) are manually assigned descriptors. The goal is to determine the values of the network parameters so that the function computed by the network approximates the function \(g\), even on inputs not in the training data that have never been seen before.
If the function computed by the network approximates \(g\) only for the training data and doesn’t approximate it well for data not in the training set, then overfitting may have occurred. This can happen if noise or random fluctuations in the training data are learnt by the network. If such inessential features are disproportionately present in the training data, then the network may fail to learn to ignore them and so not robustly generalise from the training examples. How to ovoid overfitting is an important topic, but is not considered here.
The backpropagation algorithm implements a machine learning method called gradient descent. This iterates through the learning data calculating an update for the parameter values derived from each given argument-result pair. These updates are calculated using derivatives of the functions corresponding to the neurons making up the network. When I attempted to understand the mathematics underlying this I found I’d pretty much completely forgotten the required notations and concepts of elementary calculus, although I must have learnt them once at school. I therefore needed to review some basic mathematics, such as gradients, tangents, differentiation before trying to explain why and how the gradient descent method works.
When the updates calculated from each argument-result pair are applied to a network depends on the machine learning strategy used. Two possibilities are online learning and offline learning. These are explained very briefly below.
To illustrate how gradient descent is applied to train neural nets I’ve pinched expository ideas from the YouTube video of Lecture 12a of Winston’s MIT AI course. A toy network with four layers and one neuron per layer is introduced. This is a minimal example to show how the chain rule for derivatives is used to propagate errors backwards – i.e. backpropagation.
The analysis of the one-neuron-per-layer example is split into two phases. First, four arbitrary functions composed together in sequence are considered. This is sufficient to show how the chain rule is used. Second, the functions are instantiated to be simple neuron models (sigmoid function, weights, biases etc). With this instantiation, the form of the backpropagation calculations of updates to the neuron weight and bias parameters emerges.
Next a network is considered that still has just four layers, but now with two neurons per layer. This example enables vectors and matrices to be introduced. Because the example is so small – just eight neurons in total – it’s feasible (though tedious) to present all the calculations explicitly; this makes it clear how they can be vectorised.
Finally, using notation and expository ideas from Chapter 2 of Nielsen’s book, the treatment is generalised to nets with arbitrary numbers of layers and neurons per layer. This is very straightforward as the presentation of the two-neuron-per-layer example was tuned to make this generalisation smooth and obvious.
The behaviour of a neuron is modelled using a function and the behaviour of a neural network is got by combining the functions corresponding to the behaviours of individual neurons it contains. This section outlines informally some \(\lambda\)-calculus and functional programming notation used to represent these functions and their composition.
If \(E\) is an expression containing a variable \(x\) (e.g. \(E = x^2\)), then \(\lambda x. E\) denotes the function that when applied to an argument \(a\) returns the value obtained by evaluating \(E\) after \(a\) has been substituted for \(x\) (e.g. \(\lambda x. x^2\) denotes the squaring function \(a \mapsto a^2\)). The notation \(E[a/x]\) is commonly used for the expression resulting from substituting \(a\) for \(x\) in \(E\) (e.g. \(x^2[a/x] = a^2\)).
The usual notation for the application of a function \(\phi\) to arguments \(a_1\), \(a_2\), \(\ldots\), \(a_n\) is \(\phi(a_1,a_2,\ldots,a_n)\). Although function applications are sometimes written this way here, the notation \(\phi~a_1~a_2~\ldots~a_n\), where the brackets are omitted, is also often used. The reason for this is because multi-argument functions are often conveniently represented as single argument functions that return functions, using a trick called “currying” elaborated further below. For example, the two-argument addition function can be represented as the one-argument function \(\lambda x. \lambda y. x + y\). Such functions are applied to their arguments one at a time. For example
\((\lambda x. \lambda y. x + y) 2 = (\lambda y. x + y)[2/x] = (\lambda y. 2 + y)\)
and then the resulting function can be applied to a second argument
\((\lambda y. 2 + y) 3 = 2 + 3 = 5\)
and so \(((\lambda x. \lambda y. x + y) 2) 3 = (\lambda y. 2 + y) 3 = 5\).
Functions of the form \(\lambda x. \lambda y. E\) can be abbreviated to \(\lambda x\hskip1mm y. E\), e.g. \(\lambda x\hskip1mm y. x+y\). More generally \(\lambda x_1. \lambda x_2.~\cdots~\lambda x_n. E\) can be abbreviated to \(\lambda x_1\hskip1mm x_2\hskip1mm \cdots\hskip1mm x_n. E\).
By convention, function application associates to the left, i.e. \(\phi~a_1~a_2~\ldots~a_{n-1}~a_n\) is read as \(((\cdots((\phi~a_1)~a_2)~\ldots~a_{n-1})~a_n)\). For example, \(((\lambda x{\hskip1mm} y. x + y) 2) 3\) can be written as \((\lambda x{\hskip1mm} y. x + y) 2\hskip1mm 3\).
Functions that take their arguments one-at-a-time are called curried. The conversion of multi-argument functions to curried functions is called currying. Applying a curried function to fewer arguments than it needs is called partial application, for example \((\lambda x{\hskip1mm} y. x + y) 2\) is a partial application. The result of such an application is a function which can then be applied to the remaining arguments, as illustrated above. Note that, in my opinion, the Wikipedia page on partial application is rather confused and unconventional.
Functions that take pairs of arguments or, more generally, take tuples or vectors as arguments, can be expressed as \(\lambda\)-expressions with the notation \(\lambda(x_1,\ldots,x_n). E\); pairs being the case when \(n=2\). For example, \(\lambda(x,y). x+y\) is a way of denoting a non-curried version of addition, where:
\((\lambda (x,y). x + y) (2,3) = 2 + 3 = 5\)
The function \(\lambda(x,y). x+y\) only makes sense if it is applied to a pair. Care is needed to make sure curried functions are applied to arguments ‘one at a time’ and non-curried functions are applied to tuples of the correct length. If this is not done, then either meaningless expressions result like \((\lambda (x,y). x + y)\hskip1mm 2\hskip1mm 3\) where \(2\) is not a pair or \((\lambda x{\hskip1mm} y. x + y) (2,3) = \lambda y. (2,3) + y\) where a pair is substituted for \(x\) resulting in nonsense.
The notation \(\lambda(x_1,\ldots,x_n). E\) is not part of a traditional logician’s standard version of the \(\lambda\)-calculus, in which only single variables are bound using \(\lambda\), but it is common in functional programming.
A statement like \(\phi : {\Bbb R} \rightarrow {\Bbb R}\) is sometimes called a type judgement. It means that \(\phi\) is a function that takes a single real number as an argument and returns a single real number as a result. For example: \(\lambda x. x^2~:~{\Bbb R} \rightarrow {\Bbb R}\). More generally, \(\phi : \tau_1 \rightarrow \tau_2\) means that \(\phi\) takes an argument of type \(\tau_1\) and returns a result of \(\tau_2\). Even more generally, \(E : \tau\) means that \(E\) has the type \(\tau\). The kind of type expressions that \(\tau\) ranges over are illustrated with examples below.
The type \(\tau_1\rightarrow\tau_2\) is the type of functions that take an argument of type \(\tau_1\) and return a result of type \(\tau_2\). By convention, \(\rightarrow\) to associates to the right: read \(\tau_1\rightarrow\tau_2\rightarrow\tau_3\) as \(\tau_1\rightarrow(\tau_2\rightarrow\tau_3)\). The type \(\tau_1\times\tau_2\) is the type of pairs \((t_1,t_2)\), where \(t_1\) has type \(\tau_1\) and \(t_2\) has type \(\tau_2\), i.e \(t_1:\tau_1\) and \(t_2:\tau_2\). Note that \(\times\) is more tightly binding than \(\rightarrow\), so read \(\tau_1 \times \tau_2 \rightarrow \tau_3\) as \((\tau_1 \times \tau_2) \rightarrow \tau_3\), not as \(\tau_1 \times (\tau_2 \rightarrow \tau_3)\). The type \(\tau^n\) is the type of \(n\)-tuples of values of type \(\tau\), so \({\Bbb R}^2 = {\Bbb R} \times {\Bbb R}\). I hope the following examples are sufficient to make the types used here intelligible; \(D\) and \(\nabla\) are explained later.
\(\begin{array}{lll} 2 : {\Bbb R} & \lambda x. x^2 : {\Bbb R} \rightarrow {\Bbb R} & (\lambda x. x^2)\hskip1mm 2 : {\Bbb R} \\ \lambda x{\hskip1mm} y. x {+} y : {\Bbb R} \rightarrow {\Bbb R} \rightarrow {\Bbb R} & (\lambda x{\hskip1mm} y. x {+} y) 2: {\Bbb R} \rightarrow {\Bbb R} & (\lambda x{\hskip1mm} y. x {+} y)\hskip1mm 2\hskip1mm 3 : {\Bbb R} \\ (2,3) : {\Bbb R} \times {\Bbb R} & \lambda(x,y). x{+}y : {\Bbb R} \times {\Bbb R} \rightarrow {\Bbb R} & (\lambda (x,y). x {+} y) (2,3): {\Bbb R} \\ D : ({\Bbb R} \rightarrow {\Bbb R}) \rightarrow ({\Bbb R} \rightarrow {\Bbb R}) & \nabla : ({\Bbb R}^n\rightarrow{\Bbb R}) \rightarrow ({\Bbb R}\rightarrow{\Bbb R})^n \\ \end{array}\)
A two-argument function that is written as an infix – i.e. between its arguments – is called an operator. For example, \(+\) and \(\times\) are operators that correspond to the addition and multiplication functions. Sometimes, to emphasise that an operator is just a function, it is written as a prefix, e.g. \(x+y\) and \(x\times y\) are written as \(+~x~y\) and \(\times~x~y\). This is not done here, but note that the product of numbers \(x\) and \(y\) will sometimes be written with an explicit multiplication symbol as \(x \times y\) and sometimes just as \(x\hskip1mm y\).
The types of operators are shown by ignoring their infix aspect, for example: \(+ : {\Bbb R} \times {\Bbb R} \rightarrow {\Bbb R}\) and \(\times : {\Bbb R} \times {\Bbb R} \rightarrow {\Bbb R}\) (the two occurrences of the symbol “\(\times\)” in the immediately preceding type judgement for \(\times\) stand for different things: the leftmost is the multiplication operator and the other one is type notation for pairing). I’ve given \(+\) and \(\times\) non-curried types as this is more in line with the usual way of thinking about them in arithmetic, but giving them the type \({\Bbb R} \rightarrow {\Bbb R} \rightarrow {\Bbb R}\) would also make sense.
An important operator is function composition denoted by “\(\circ\)” and defined by \((\phi_1 \circ \phi_2)(x) = \phi_1(\phi_2~x)\) or alternatively \(\circ = \lambda (\phi_1,\phi_2). \lambda x. \phi_1(\phi_2~x)\). The type of function composition is \(\circ : ((\tau_2\rightarrow\tau_3) \times (\tau_1\rightarrow\tau_2)) \rightarrow (\tau_1\rightarrow\tau_3)\), where \(\tau_1\), \(\tau_2\), \(\tau_3\) can be any types.
The ad hoc operator \({\star}: ({\Bbb R}\rightarrow{\Bbb R})^n \times {\Bbb R}^n \rightarrow {\Bbb R}^n\) is used here to apply a tuple of functions pointwise to a tuple of arguments and return the tuple of results: \((\phi_1,\ldots,\phi_n)\;{\star}\; (p_1,\ldots,p_n) = (\phi_1~p_1,\ldots,\phi_n~p_n)\). I arbitrarily chose \({\star}\) as I couldn’t find a standard symbol for such pointwise applications of vectors of functions.
By convention, function application associates to the left, so \(\phi_1~\phi_2~a\) means \((\phi_1~\phi_2)~a\), not \(\phi_1(\phi_2~a)\). An example that comes up later is \(\nabla\phi(p_1,p_2)\), which by left association should mean \((\nabla\phi)(p_1,p_2)\). In fact \((\nabla\phi)(p_1,p_2)\) doesn’t make sense – it’s not well-typed – but it’s useful to give it the special meaning of being an abbreviation for \((\nabla\phi)\;{\star}\;(p_1,p_2)\); this meaning is a one-off notational convention for \(\nabla\). For more on \(\nabla\) see the section entitled The chain rule below.
The goal of the kind of machine learning described here is to use it to train a network to approximate a given function \(g\). The training aims to discover a value for a network parameter \(p\), so that with this parameter value the behaviour of the network approximates \(g\). The network is denoted by \({\small\textsf{N}}(p)\) to make explicit that it has the parameter \(p\). The behaviour of \({\small\textsf{N}}(p)\) is represented by a function \(f_{\scriptsize\textsf{N}}~p\). Thus the goal is to discover a value for \(p\) so that \(g\) is approximated by \(f_{\scriptsize\textsf{N}}~p\), i.e. \(g~x\) is close to \(f_{\scriptsize\textsf{N}}~p~x\) for those inputs \(x\) in the training set (which hopefully are typical of the inputs the network is intended to be used on).
Note that \(f_{\scriptsize\textsf{N}}\) is a curried function taking two arguments. If it is partially applied to its first argument then the result is a function expecting the second argument: \(f_{\scriptsize\textsf{N}}~p\) is a function and \(f_{\scriptsize\textsf{N}}~p~x\) is this function applied to \(x\).
The goal is thus to learn a value for \(p\) so \(f_{\scriptsize\textsf{N}}~p~x\) and \(g~x\) are close for values of \(x\) in the learning set. The parameter \(p\) for an actual neural network is a vector of real numbers consisting of weights and biases, but for the time being this is abstracted away and \(p\) is simplified to a single real number. Weights and biases appear in the section entitled Neuron models and cost functions below.
The value \(p\) is learnt by starting with a random initial value \(p_0\) and then training with a given set of input-output examples \(\{(x_1,~g~x_1),(x_2,~g~x_2),(x_3,~g~x_3), \ldots\}\). There are two methods for doing this training: online learning (also called one-step learning) and offline learning (also called batch learning). Both methods first compute changes, called deltas, to \(p\) for each training pair \((x_i,~g~x_i)\).
Online learning consists in applying a delta to the parameter after each example. Thus at the \(i^{\textrm{th}}\) step of training, if \(p_i\) is the parameter value computed so far, then \(g~x_i\) and \(f_{\scriptsize\textsf{N}}~p_i~x_i\) are compared and a delta, say \({\Delta}p_i\), is computed, so \(p_{i{+}1} = p_i + {\Delta}p_i\) is the parameter value used in the \(i{+}1^{\textrm{th}}\) step.
Offline learning consists in first computing the deltas for all the examples in the given set, then averaging them, and then finally applying the averaged delta to \(p_0\). Thus if there are \(n\) examples and \(p_i\), \({\Delta}p_i\) are the parameter value and the delta computed at the \(i^{\textrm{th}}\) step, respectively, then \(p_i=p_0\) for \(i<n\) and \(p_n = p_0 + (\frac{1}{n} \times \Sigma_{i{=}1}^{i{=}n} {\Delta}p_i)\).
If there are \(n\) examples, then with online learning \(p\) is updated \(n\) times, but with offline learning it is only updated once.
Apparently online learning converges faster, but offline learning results in a network that makes fewer errors. With offline learning, all the examples can be processed in parallel (e.g. on a GPU) leading to faster processing. Online learning is inherently sequential.
In practice offline learning is done in batches, called learning epochs. A subset of the given examples is chosen ‘randomly’ and then processed as above. Another subset is then chosen from the remaining unused training examples and processed, and so on until all the examples have been processed. Thus the learning computation is a sequence of epochs of the form ‘choose-new-subset-then-update-parameter’ which are repeated until all the examples have been used.
The computation of the deltas is done by using the structure of the network to precompute the way changes in \(p\) effect the function \(f_{\scriptsize\textsf{N}}~p\), so for each training pair \((x,~g~x)\) the difference between the desired output \(g~x\) and the output of the network \(f_{\scriptsize\textsf{N}}~p~x\) can be used to compute a delta that when applied to \(p\) reduces the error.
The backpropagation algorithm is an efficient way of computing such parameter changes for a machine learning method called gradient descent. Each training example \((x_i,~g~x_i)\) is used to calculate a delta to the parameter value \(p\). With online learning the changes are applied after processing each example; with offline learning they are first computed for the current epoch and they are then averaged and applied.
Gradient descent is an iterative method for finding the minimum of a function. The function to be minimised is called the cost function and measures the closeness of a desired output \(g~x\) for an input \(x\) to the output of the network, i.e. \(f_{\scriptsize\textsf{N}}~p~x\).
Suppose \(C\) is the cost function, so \(C~y~z\) measures the closeness of a desired output \(y\) to the network output \(z\). The goal is then is to learn a value \(p\) that minimises \(C(g~x)(f_{\scriptsize\textsf{N}}~p~x)\) for values of \(x\) important for the application.
At the \(i^{\textrm{th}}\) step of gradient descent one evaluates \(C(g~x_i)(f_{\scriptsize\textsf{N}}~p_i~x_i)\) and uses the yet-to-be-described backpropagation algorithm to determine a delta \({\Delta}p_i\) to \(p\) so \(p_{i+1} = p_i + {\Delta}p_i\).
Gradient descent is based on the fact that \(\phi~a\) decreases fastest if one changes \(a\) in the direction of the negative gradient of the tangent of the function at \(a\).
The next few sections focus on a single step, where the input is fixed to be \(x\), and explain how to compute \({\Delta}p\) so that \(\phi(p) > \phi(p{+}{\Delta}p)\), where \(\phi~p = C(g~x)(f_{\scriptsize\textsf{N}}~p~x)\).
The tangent of a curve is the line that touches the curve at a point and ‘is parallel’ to the curve at that point. See the red tangents to the black curve in the diagram below.

The gradient of a line is computed by taking any two distinct points on it and dividing the change in y-coordinate between the two points divided by the corresponding change in x-coordinate.
Consider the points \((2,0)\) an \((4,2)\) on the red tangent line to the right of the diagram above: the y coordinate changes from \(0\) to \(2\), i.e. by \(2\), and the x coordinate changes from \(2\) to \(4\), i.e. also by \(2\). The gradient is thus \(2/2 = 1\) and hence so the negative gradient is \(-1\).
Consider now the points \((-3,1)\) an \((-1,0)\) on the red tangent line to the left of the diagram: the y coordinate changes from \(0\) to \(1\), i.e. by \(1\), and the x coordinate changes from \(-1\) to \(-3\), i.e. by \(-2\). The gradient is thus \(1/-2 = -0.5\) and hence the negative gradient is \(0.5\).
This example is for a function \({\phi}:{\Bbb R}\rightarrow{\Bbb R}\) and the direction of the tangent is just whether its gradient is positive or negative. The diagram illustrates that for a sufficiently small change to the argument of \({\phi}\) in the direction of the negative gradient decreases the value of \({\phi}\), i.e. for a sufficiently small \(\eta>0\), \({\phi}(x-\eta{\times}\textrm{gradient-at-}x)\) is less than \({\phi}(x)\). Thus, if \(dx = -\eta{\times}\textrm{gradient-at-}x\) then \({\phi}(x{+}dx)\) is less than \({\phi}(x)\).
The gradient at \(p\) of \({\phi}:{\Bbb R}\rightarrow{\Bbb R}\) is given by value of the derivative of \(\phi\) at \(p\). The derivative function of \({\phi}\) is denoted by \(D\phi\) in Euler’s notation.
Other notations are: \(\phi'\) due to Lagrange, \(\dot{\phi}\) due to Newton and \(\frac{d\phi}{dx}\) due to Leibniz.
Using Euler notation, the derivative of \(\phi\) at \(p\) is \(D\phi(p)\), where \(D\) is a higher-order function \(D : ({\Bbb R}{\rightarrow}{\Bbb R})\rightarrow({\Bbb R}{\rightarrow}{\Bbb R})\).
If a function is ‘sufficiently smooth’ so that it has a derivative at \(p\), then it is called differentiable at \(p\). If \({\phi}:{\Bbb R}\rightarrow{\Bbb R}\) is differentiable at \(p\), then \({\phi}\) can be approximated by its tangent for a sufficiently small interval around \(p\). This tangent is the straight line represented by the function, \(\hat{\phi}\) say, defined by \(\hat{{\phi}}(v) = r\hskip1mm v + s\), where \(r\) is the gradient of \(\phi\) at \(p\), \(D {\phi} (p)\) in Euler notation, and \(s\) is the value that makes \(\hat{{\phi}}(p) = {\phi}(p)\), which is easily calculated:
\(\begin{array}{ll} \hat{{\phi}}(p) = {\phi}(p) & \Leftrightarrow ~~~ r\hskip1mm p + s = {\phi}(p) \\ & \Leftrightarrow ~~~ D{\phi}(p)\hskip1mm p + s = {\phi}(p) \\ & \Leftrightarrow ~~~ s = {\phi}(p) - p\hskip1mm D{\phi}(p) \end{array}\)
Thus the tangent function at \(p\) is defined by
\(\begin{array}{ll} \hat{{\phi}}(v) & = ~~~ D{\phi}(p)\hskip1mm v + {\phi}(p) - p\hskip1mm D{\phi}(p) \\ & = ~~~ {\phi}(p) + D{\phi}(p)\hskip1mm (v - p) \\ \end{array}\)
If the argument of \(\hat{{\phi}}\) is changed from \(p\) to \(p{+}{\Delta}p\), then the corresponding change to \(\hat{{\phi}}\) is easily calculated.
\(\begin{array}{ll} \hat{{\phi}}(p) - \hat{{\phi}}(p{+}{\Delta}p) & = ~~~ (r\hskip1mm p + s) - (r\hskip1mm (p{+}{\Delta}p) + s) \\ & = ~~~ r\hskip1mm p + s - r\hskip1mm (p{+}{\Delta}p) - s \\ & = ~~~ r\hskip1mm p + s - r\hskip1mm p - r\hskip1mm {\Delta}p - s \\ & = ~~~ - r\hskip1mm {\Delta}p \\ & = ~~~ - D{\phi}(p)\hskip1mm {\Delta}p \\ \end{array}\)
This illustrates that the amount \(\hat{{\phi}}(p)\) changes when \(p\) is changed depends on \(D{\phi}(p)\), i.e. the gradient of \(\phi\) at \(p\).
If \(\eta > 0\) and \({\Delta}p\) is taken to be \(-\eta\times\textrm{gradient-at-}p\) i.e. \({\Delta}p = - \eta \hskip1mmD{\phi}(p)\) then
\(\begin{array}{ll} \hat{{\phi}}(p) - \hat{{\phi}}(p{+}{\Delta}p) & = ~~~ - D{\phi}(p)\hskip1mm {\Delta}p \\ \ & = ~~~ - D{\phi}(p) (-\eta \hskip1mm D{\phi}(p)) \\ & = ~~~ \eta (D{\phi}(p))^2 \end{array}\)
and thus it is guaranteed that \(\hat{{\phi}}(p) > \hat{{\phi}}(p{+}{\Delta}p)\), therefore adding \({\Delta}p\) to the argument of \(\hat{{\phi}}\) decreases it. If \({\Delta}p\) is small enough then \({\phi}(p{+}{\Delta}p)~\approx~\hat{{\phi}}(p{+}{\Delta}p)\), where the symbol “\(\approx\)” means “is close to”, and hence
\(\begin{array}{ll} {\phi}(p) - {\phi}(p{+}{\Delta}p) &=~~~ \hat{{\phi}}(p) - {\phi}(p{+}{\Delta}p)\\ &\approx~~~ \hat{{\phi}}(p) - \hat{{\phi}}(p{+}{\Delta}p)\\ &=~~~\eta \hskip1mm D \phi (p) \cdot D \phi (p) \end{array}\)
Thus \({\Delta}p = -\eta \hskip1mm D{\phi}(p)\) is a plausible choice for the delta, so changing \(p\) to \(p{+}{\Delta}p\), i.e. to \(p - \eta\hskip1mm D{\phi}(p)\) is a plausible way to change \(p\) at each step of gradient descent. The choice of \(\eta\) determines the learning behaviour: too small makes learning slow, too large makes learning thrash and maybe even not converge.
In practice the parameter to be learnt won’t be a single real number but a vector of many, say \(k\), reals (one or more for each neuron), so \({\phi}:{\Bbb R}^k\rightarrow{\Bbb R}\). Thus gradient descent needs to be applied to a \(k\)-dimensional surface.
One step of gradient descent consists in computing a change \({\Delta}p\) to \(p\) to reduce \(C(g~x)(f_{\scriptsize\textsf{N}}~p~x)\), i.e. the difference between the correct output \(g~x\) from the training set and the output \(f_{\scriptsize\textsf{N}}~p~x\) from the network that’s being trained.
In the two-dimensional case \(k=2\) and \(p = (v_1,v_2)\) where \(v_1\), \(v_2\) are real numbers. To make a step in this case one computes changes \({\Delta}v_1\) to \(v_1\) and \({\Delta}v_2\) to \(v_2\), and then \({\Delta}p = ({\Delta}v_1,{\Delta}v_2)\) and so \(p\) is changed to \(p + {\Delta}p = (v_1{+}{\Delta}v_1, v_2{+}{\Delta}v_2)\), where the “\(+\)” in “\(p + {\Delta}p\)” is vector addition. Hopefully \(C(g~x)(f_{\scriptsize\textsf{N}}~(p+{\Delta}p)~x)\) becomes smaller than \(C(g~x)(f_{\scriptsize\textsf{N}}~p~x)\).
If \({\phi}:{\Bbb R}^2\rightarrow{\Bbb R}\) is differentiable at \(p = (v_1, v_2)\) then for a small neighbourhood around \(p\) it can be approximated by the tangent plane at \(p\), which is the linear function \(\hat{\phi}\) defined by \(\hat{{\phi}}(v_1, v_2) = r_1\hskip1mm v_1 + r_2\hskip1mm v_2 + s\), where the constants \(r_1\) and \(r_2\) are the partial derivatives of \(\phi\) that define the gradient of its tangent plane. Using the gradient operator \(\nabla\) (also called “del”), the partial derivatives \(r_1\) and \(r_2\) are given by \((r_1,r_2)= \nabla \phi (p)\). The operator \(\nabla\) is discussed in more detail in the section entitled The chain rule below.
The effect on the value of \(\hat{{\phi}}\) resulting from a change in the argument can then be calculated in the same was as before.
\(\begin{array}{ll} \hat{{\phi}}(p) - \hat{{\phi}}(p{+}{\Delta}p) & = ~~~ \hat{{\phi}}(v_1, v_2) - \hat{{\phi}}(v_1{+}{\Delta}v_1, v_2{+}{\Delta}v_2)\\ & = ~~~ (r_1\hskip1mm v_1 + r_2\hskip1mm v_2 + s) - (r_1\hskip1mm (v_1{+}{\Delta}v_1) + r_2\hskip1mm (v_2{+}{\Delta}v_2) + s) \\ & = ~~~ r_1\hskip1mm v_1 + r_2\hskip1mm v_2 + s - r_1\hskip1mm (v_1{+}{\Delta}v_1) - r_2\hskip1mm (v_2{+}{\Delta}v_2) - s \\ & = ~~~ r_1\hskip1mm v_1 + r_2\hskip1mm v_2 + s - r_1\hskip1mm (v_1{+}{\Delta}v_1) - r_2\hskip1mm (v_2{+}{\Delta}v_2) - s \\ & = ~~~ r_1\hskip1mm v_1 + r_2\hskip1mm v_2 + s - r_1\hskip1mm v_1 - r_1\hskip1mm {\Delta}v_1 - r_2\hskip1mm v_2 - r_2\hskip1mm {\Delta}v_2 - s \\ & = ~~~ - r_1\hskip1mm {\Delta}v_1 - r_2\hskip1mm {\Delta}v_2 \\ & = ~~~ - (r_1\hskip1mm {\Delta}v_1 + r_2\hskip1mm {\Delta}v_2)\\ & = ~~~ - (r_1, r_2) \cdot ({\Delta}v_1 , {\Delta}v_2) \\ & = ~~~ - \nabla \phi (p) \cdot {\Delta}p \\ \end{array}\)
The symbol “\(\cdot\)” here is the dot product of vectors. For any two vectors with the same number of components – \(k\) below – the dot product is defined by:
\((x_1,x_2,\ldots,x_m)\cdot(y_1,y_2,\ldots,y_k) = x_1\hskip1mm y_1+x_2\hskip1mm y_2+\cdots+ x_m\hskip1mm y_k\)
If \(\eta > 0\) and \({\Delta}p\) is taken to be \(-\eta\times\textrm{gradient-vector-at-}p\) i.e. \({\Delta}p = -\eta \hskip1mm \nabla \phi (p)\) then
\(\begin{array}{ll} \hat{{\phi}}(p) - \hat{{\phi}}(p{+}{\Delta}p) & = ~~~ - \nabla \phi (p) \cdot {\Delta}p \\ & = ~~~ - \nabla \phi (p) \cdot (-\eta \hskip1mm \nabla \phi p) \\ & = ~~~ \eta \hskip1mm \nabla \phi (p) \cdot \nabla \phi (p) \\ \end{array}\)
Since \(\nabla \phi (p) \cdot \nabla \phi (p) = (r_1+r_2) \cdot (r_1+r_2) = r_1^2+r_2^2\), it’s guaranteed \(\hat{{\phi}}(p) > \hat{{\phi}}(p{+}{\Delta}p)\), therefore adding \({\Delta}p\) to the argument of \(\hat{{\phi}}\) decreases it. Just like in the one-dimensional case, if \({\Delta}p\) is small enough, then \({\phi}(p{+}{\Delta}p)~\approx~\hat{{\phi}}(p{+}{\Delta}p)\) and so
\(\begin{array}{ll} {\phi}(p) - {\phi}(p{+}{\Delta}p) &=~~~ \hat{{\phi}}(p) - {\phi}(p{+}{\Delta}p)\\ &\approx~~~ \hat{{\phi}}(p) - \hat{{\phi}}(p{+}{\Delta}p)\\ &=~~~\eta \hskip1mm \nabla \phi (p) \cdot \nabla \phi (p) \end{array}\)
Thus \({\Delta}p = - \eta \hskip1mm \nabla \phi (p)\), for some \(\eta>0\), is a plausible delta, so changing \(p\) to \(p{+}{\Delta}p\), i.e. to \(p - \eta\hskip1mm\nabla{\phi} (p)\), is a plausible way to change \(p\) at each step of gradient descent.
Note that \({\Delta}p = - \eta \hskip1mm \nabla \phi (p)\) is an equation between pairs. If \({\Delta}p = (r_1,r_2)\) then the equation is \(({\Delta}p_1, {\Delta}p_2) = (-\eta\hskip1mm r_1,~-\eta\hskip1mm r_2)\), which means \({\Delta}p_1 = -\eta\hskip1mm r_1\) and \({\Delta}p_2 = -\eta\hskip1mm r_2\).
The analysis when \({\phi}:{\Bbb R}^k\rightarrow{\Bbb R}\) is a straightforward generalisation of the \(k=2\) case just considered. As before, one step of gradient descent consists in computing a change \({\Delta}p\) to \(p\) to reduce \(C(g~x)(f_{\scriptsize\textsf{N}}~p~x)\), i.e. the difference between the correct output \(g~x\) from the training set and the output \(f_{\scriptsize\textsf{N}}~p~x\) from the network that’s being trained.
Replaying the \(k=2\) analysis for arbitrary \(k\) results in the conclusion that \({\Delta}p = - \eta \hskip1mm \nabla \phi (p)\) is a plausible choice of a delta for machine learning. Here, the parameter \(p\) is a \(k\)-dimensional vector of real numbers and \({\Delta}p\) is a \(k\)-dimensional vector of deltas, also real numbers.
Euler notation for derivatives is natural if one is familiar with functional programming or the \(\lambda\)-calculus. Differentiation is the application of a higher-order function \(D\) and partial derivatives are defined using \(\lambda\)-notation. Unfortunately, although this notation is logically perspicuous, it can be a bit clunky for calculating derivatives of particular functions. For such concrete calculations, the more standard Leibniz notation can work better. This is now reviewed.
The Euler differentiation operator is a function \(D : ({\Bbb R}{\rightarrow}{\Bbb R}) \rightarrow ({\Bbb R}{\rightarrow}{\Bbb R})\), but what does the Leibniz notation \(\frac{dy}{dx}\) denote?
Leibnitz notation is based on a view in which one has a number of variables, some dependent on others, with the dependencies expressed using equations like, for example, \(y = x^2\). Here a variable \(y\) is dependent on a variable \(x\) and to emphasise this one sometimes writes \(y(x) = x^2\). When using Leibnitz notation one thinks in terms of equations rather than ‘first class functions’ like \(\lambda x. x^2\).
In Leibniz notation the derivative of \(y\) with respect to \(x\) is denoted by \(\frac{dy}{dx}\).
From the equation \(y=x^2\) one can derive the equation \(\frac{dy}{dx} = 2 \times x\) by using rules for differentiation.
In this use case, \(\frac{dy}{dx}\) is a variable dependent on \(x\) satisfying the derived equation. This is similar to Newton’s notation \(\dot{y}\), where the variable \(y\) depends on is taken from the equation defining \(y\).
The expression \(\frac{dy}{dx}\) is also sometimes used to denote the function \(D(\lambda x. x^2) = \lambda x. 2x\mathrm{.}\) In this use case one can write \(\frac{dy}{dx}(a)\) to mean the derivative evaluated at point \(a\) - i.e. \(D(\lambda x. x^2)(a)\).
It would be semantically clearer to write \(y(x) = x^2\) and \(\frac{dy}{dx}(x) = 2 \times x\), but this is verbose and so “\((x)\)” is omitted and the dependency on \(x\) inferred from context.
Sometimes \(\left.\frac{dy}{dx}\right\rvert_{x=a}\) is written instead of \(\frac{dy}{dx}(a)\).
I find this use of \(\frac{dy}{dx}\) as both a variable and a function rather confusing.
Yet another Leibniz style notation is to use \(\frac{d}{dx}\) instead of \(D\). One then writes \(\frac{d}{dx}(f)\) or \(\frac{d(f)}{dx}\) to mean \(Df\). If there is only one variable \(x\) being varied then the \(dx\) is redundant, but the corresponding notation for partial derivatives is more useful.
The derivative \(\frac{dy}{dx}\) assumes \(y\) depends on just one variable \(x\). If \(y\) depends on more than one variable, for example \(y = \sin(x) + z^2\), where \(y\) depends on \(x\) and \(z\), then the partial derivatives \(\frac{\partial y}{\partial x}\) and \(\frac{\partial y}{\partial z}\) are the Leibniz notations for the partial derivatives with respect to \(y\) and \(z\).
The partial derivative with respect to a variable is the ordinary derivative with respect to the variable, but treating the other variables as constants.
If \(\phi : {\Bbb R}^k\rightarrow{\Bbb R}\), then using Euler notation the partial derivatives are \(D(\lambda v_i. \phi(v_1,\ldots,v_i,\ldots,v_k))\) for \(1 \leq i \leq k\).
For example, if \(\phi(x,z) = \sin(x) + z^2\) – i.e. \(\phi = \lambda(x,z). \sin(x) + z^2\) – then the partial derivatives are \(D(\lambda x. \sin(x) + z^2))\) and \(D(\lambda z. \sin(x) + z^2))\).
The expression \(\frac{\partial(\phi(v_1,\ldots,v_i,\ldots,v_k))}{\partial v_i}\) is the Leibniz notation for the \(i^{\textrm{th}}\) partial derivative of the expression \(\phi(v_1,\ldots,v_i,\ldots,v_k)\).
If \(v\) is defined by the equation \(v = \phi(v_1,\ldots,v_k)\), then the Leibniz notation for the \(i^{\textrm{th}}\) partial derivatives is \(\frac{\partial v}{\partial v_i}\).
The partial derivative with respect to a variable is calculated using the ordinary rules for differentiation, but treating the other variables as constants.
For example, if \(y = \sin(x) + z^2\) then by the addition rule for differentiation:
\(\begin{array}{l} \frac{\partial y}{\partial x} = \frac{\partial(\sin(x))}{\partial x} + \frac{\partial(z^2)}{\partial x} \\ \phantom{\frac{\partial y}{\partial x}} = \cos(x) + 0 \end{array}\)
This is so because \(\frac{d(\sin(x))}{dx} = \cos(x)\) and if \(z\) is being considered as a constant, then \(z^2\) is also constant, so its derivative is \(0\).
The partial derivative with respect to \(z\) is
\(\begin{array}{l} \frac{\partial y}{\partial z} = \frac{\partial(\sin(x))}{\partial z} + \frac{\partial(z^2)}{\partial z} \\ \phantom{\frac{\partial y}{\partial z}} = 0 + 2 \times z \end{array}\)
because \(\frac{\partial(\sin(x))}{\partial z}\) is \(0\) if \(x\) is considered as a constant and \(\frac{\partial(z^2)}{\partial z} = \frac{d(z^2)}{dz} = 2 \times z\).
A network processes an input by a sequence of ‘layers’. Here is an example from Chapter 1 of Nielsen’s book.
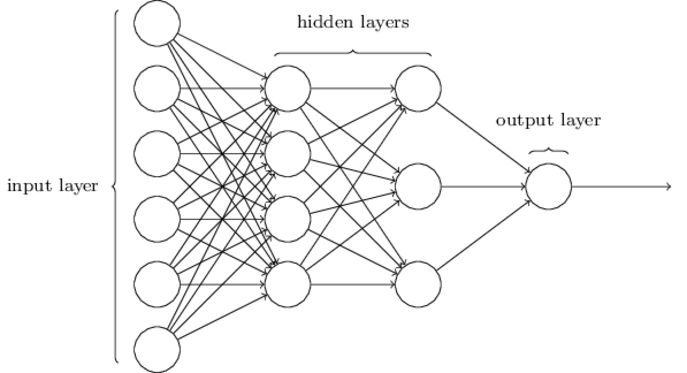
\(\hskip 39mm f_{\scriptsize L1} : {\Bbb R}^6 {\rightarrow} {\Bbb R}^6\) \(\hskip 5mm f_{\scriptsize L2} : {\Bbb R}^6 {\rightarrow} {\Bbb R}^4\) \(\hskip 5mm f_{\scriptsize L3} : {\Bbb R}^4 {\rightarrow} {\Bbb R}^3\) \(\hskip 5mm f_{\scriptsize L4} : {\Bbb R}^3 {\rightarrow} {\Bbb R}^1\)
This network has four layers: an input layer, two hidden layers and an output layer. The input of each layer, except for the input layer, is the output of the preceding layer. The function computed by the whole network is the composition of functions corresponding to each layer \(f_{\scriptsize L4} \circ f_{\scriptsize L3} \circ f_{\scriptsize L2} \circ f_{\scriptsize L1}\). Here \(f_{\scriptsize L1}\) is the function computed by the input layer, \(f_{\scriptsize L2}\) the function computed by the first hidden layer, \(f_{\scriptsize L3}\) the function computed by the second hidden layer, and \(f_{\scriptsize L4}\) the function computed by the output layer. Their types are shown above, where each function is below the layer it represents.
By the definition of function composition “\(\circ\)”
\((f_{\scriptsize L4} \circ f_{\scriptsize L3} \circ f_{\scriptsize L2} \circ f_{\scriptsize L1})~x = f_{\scriptsize L4}(f_{\scriptsize L3}(f_{\scriptsize L2}(f_{\scriptsize L1}~x)))\)
The circular nodes in the diagram represent neurons. Usually each neuron has two parameters: a weight and a bias, both real numbers. In some treatments, extra constant-function nodes are added to represent the biases, and then all nodes just have a single weight parameter. This bias representation detail isn’t important here (it is considered in the section entitled Neuron models and cost functions below). In the example, nodes have both weight and bias parameters, so the function computed by a layer is parametrised on \(2\times\) the number of neurons in the layer. For the example network there are \(2 \times 14\) parameters in total. It is the values of these that the learning algorithm learns. To explain backpropagation, the parameters need to be made explicit, so assume given functions \(f_j\) for each layer \(j\) such that \(f_{\scriptsize Lj} = f_j(p_j)\), where \(p_j\) is the vector of parameters for layer \(j\). For our example:
\(\begin{array}{l} f_1 : {\Bbb R}^{12} \rightarrow {\Bbb R}^6 \rightarrow {\Bbb R}^6 \\ f_2 : {\Bbb R}^8 \rightarrow {\Bbb R}^6 \rightarrow {\Bbb R}^4 \\ f_3 : {\Bbb R}^6 \rightarrow {\Bbb R}^4 \rightarrow {\Bbb R}^3 \\ f_4 : {\Bbb R}^2 \rightarrow {\Bbb R}^3 \rightarrow {\Bbb R}^1 \\ \end{array}\)
The notation used at the beginning of this article was \(f_{\scriptsize\textsf{N}}~p\) for the function computed by a net \({\small\textsf{N}}(p)\), \(p\) being the parameters. For the example above, \(p=(p_1,p_2,p_3,p_4)\) and \(f_{\scriptsize\textsf{N}}~p~x = f_4~p_4(f_3~p_3(f_2~p_2(f_1~p_1~x)))\).
The goal of each learning step is to compute \({\Delta}p_1\), \({\Delta}p_2\), \({\Delta}p_3\), \({\Delta}p_4\) so that if \(\phi(p_1,p_2,p_3,p_4) = C(g~x)(f_4~p_4(f_3~p_3(f_2~p_2(f_1~p_1~x))))\) then \(\phi(p_1{+}{\Delta}p_1,p_2{+}{\Delta}p_2,p_3{+}{\Delta}p_3,p_4{+}{\Delta}p_4)\) is less than \(\phi(p_1,p_2,p_3,p_4)\).
By arguments similar to those given earlier, a plausible choice of deltas is \({\Delta}p = ({\Delta}p_1,{\Delta}p_2,{\Delta}p_3,{\Delta}p_4) = - \eta \hskip1mm \nabla \phi(p)\), where – as elaborated below – \(\nabla \phi (p)\) is the vector of the partial derivatives of \(\phi\).
To see how the backpropagation algorithm calculates these backwards, it helps to first look at a linear net.
Consider the linear network shown in the diagram below
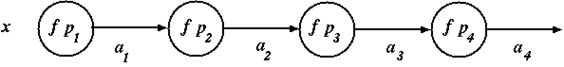
where \(a_i\) is the output from the \(i^{\textrm{th}}\) layer, called the \(i^{\textrm{th}}\) activation, defined by
\(\begin{array}{l} a_1 = f~p_1~x \\ a_2 = f~p_2~(f~p_1~x) \\ a_3 = f~p_3~(f~p_2~(f~p_1~x)) \\ a_4 = f~p_4~(f~p_3~(f~p_2~(f~p_1~x))) \\ \end{array}\)
The activation \(a_{i{+}1}\) can be defined in terms of \(a_i\).
\(\begin{array}{l} a_2 = f~p_2~a_1 \\ a_3 = f~p_3~a_2 \\ a_4 = f~p_4~a_3 \\ \end{array}\)
The input \(x\) and activations \(a_i\) are real numbers. For simplicity, bias parameters will be ignored for now, so the parameters \(p_i\) are also real numbers and thus \(f : {\Bbb R}\rightarrow{\Bbb R}\rightarrow{\Bbb R}\).
The vector equation \({\Delta}p = - \eta \hskip1mm \nabla \phi (p)\) elaborates to
\(({\Delta}p_1,{\Delta}p_2,{\Delta}p_3,{\Delta}p_4) = - \eta \hskip1mm \nabla \phi (p_1,p_2,p_3,p_4)\)
This section starts by showing why taking the derivatives of compositions of functions is needed for calculating deltas. The chain rule for differentiation is used to do this and so it is then stated and explained.
The derivative operator \(D\) maps a function \(\phi : {\Bbb R}\rightarrow{\Bbb R}\) to the function \(D\phi : {\Bbb R}\rightarrow{\Bbb R}\) such that \(D \phi(p)\) is the derivative – i.e. gradient – of \(\phi\) at \(p\). Thus \(D : ({\Bbb R}{\rightarrow}{\Bbb R}) \rightarrow ({\Bbb R}{\rightarrow}{\Bbb R})\).
When \(\phi : {\Bbb R}^k \rightarrow {\Bbb R}\) is a real-valued function of \(k\) arguments, then according to the theory of differentiation, the gradient is given by \(\nabla\phi\), where \(\nabla : ({\Bbb R}^k \rightarrow {\Bbb R}) \rightarrow ({\Bbb R}\rightarrow{\Bbb R})^k\).
\(\nabla\phi\) is a vector of functions, where each function gives the rate of change of \(\phi\) with respect to a single argument. These rates of change functions are the partial derivatives of \(\phi\) and are the derivatives when one argument is varied and the others are held constant. The derivative when all variables are allowed to vary is the total derivative; this is not used here.
If \(1 \leq i \leq k\) then the \(i^{\textrm{th}}\) partial derivative of \(\phi\) is \(D~(\lambda {\color{red}v}. \phi(p_1,\ldots,{\color{red}v},\ldots,p_k))\), where the argument being varied – the \(\lambda\)-bound variable – is highlighted in red. Thus \({\nabla}\phi = (D~(\lambda {\color{red}v}. \phi({\color{red}v},p_2,\ldots,p_k)), \ldots,\hskip1mm D~(\lambda {\color{red}v}. \phi(p_1,p_2,\ldots,{\color{red}v})))\).
Note that \(\phi : {\Bbb R}^k \rightarrow {\Bbb R}\), but \(\lambda {\color{red}v}. \phi(p_1,\ldots,{\color{red}v},\ldots,p_k) : {\Bbb R}\rightarrow{\Bbb R}\) and \(\nabla\phi : ({\Bbb R}\rightarrow{\Bbb R})^k\).
Thus if \((p_1,p_2,\ldots,p_k)\) is a vector of \(k\) real numbers, then \({\nabla}\phi(p_1,p_2,\ldots,p_k)\) is not well-typed, because by the left association of function application, \({\nabla}\phi(p_1,p_2,\ldots,p_k)\) means \(({\nabla}\phi)(p_1,p_2,\ldots,p_k)\). However, it is useful to adopt the convention that \({\nabla}\phi(p_1,p_2,\ldots,p_k)\) denotes the vector of the results of applying each partial derivative to the corresponding \(p_i\) – i.e. defining:
\(\begin{array}{ll} {\nabla}\phi(p_1,\ldots,p_i,\ldots,p_k) ~~~= & (D~(\lambda {\color{red}v}. \phi({\color{red}v},\ldots,p_i,\ldots,p_k))~p_1, \\ & \phantom{(}~ \vdots\\ & \phantom{(}D~(\lambda {\color{red}v}. \phi(p_1,\ldots,{\color{red}v},\ldots,p_k))~p_i, \\ & \phantom{(}~ \vdots\\ & \phantom{(}D~(\lambda {\color{red}v}. \phi(p_1,\ldots,p_i,\ldots,{\color{red}v}))~p_k) \end{array}\)
This convention has already been used above when \(\phi : {\Bbb R}{\times}{\Bbb R} \rightarrow {\Bbb R}\) and it was said that \(\nabla \phi(p_1,p_2) = (r_1,r_2)\), where \(r_1\) and \(r_2\) determine the gradient of the tangent plane to \(\phi\) at point \((p_1,p_2)\).
If the operator \({\star}\) is defined to apply a tuple of functions pointwise to a tuple of arguments by
\((\phi_1,\ldots,\phi_k)\;{\star}\; (p_1,\ldots,p_k) = (\phi_1~p_1,\ldots,\phi_k~p_k)\)
then \(\nabla \phi (p_1,\ldots,p_k)\) is an abbreviation for \(\nabla \phi\; {\star}\; (p_1,\ldots,p_k)\).
To calculate each \({\Delta}p_i\) for the four-neuron linear network example one must calculate \(\nabla\phi(p_1,p_2,p_3,p_4)\) since \(({\Delta}p_1,{\Delta}p_2,{\Delta}p_3,{\Delta}p_4) = - \eta \hskip1mm \nabla \phi (p_1,p_2,p_3,p_4)\) and hence \({\Delta}p_i\) is the \(i^{\textrm{th}}\) partial derivative function of \(\phi\) applied to the parameter value \(p_i\).
Recall the definition of \(\phi\)
\(\begin{array}{l} \phi(p_1,p_2,p_3,p_4) = C(g~x)(f~p_4(f~p_3(f~p_2(f~p_1~x))))\\ \end{array}\)
Hence
\(\begin{array}{l} {\Delta}p_1 = - \eta\hskip1mm D(\lambda v. C(g~x)(f~p_4(f~p_3(f~p_2(f~v~x))))\hskip1mm p_1\\ {\Delta}p_2 = - \eta\hskip1mm D(\lambda v. C(g~x)(f~p_4(f~p_3(f~v(f~p_1~x))))\hskip1mm p_2\\ {\Delta}p_3 = - \eta\hskip1mm D(\lambda v. C(g~x)(f~p_4(f~v(f~p_2(f~p_1~x))))\hskip1mm p_3\\ {\Delta}p_4 = - \eta\hskip1mm D(\lambda v. C(g~x)(f~v(f~p_3(f~p_2(f~p_1~x))))\hskip1mm p_4\\ \end{array}\)
which can be reformulated using the function composition operator “\(\circ\)” to
\(\begin{array}{l} {\Delta}p_1 = - \eta\hskip1mm D(C(g~x) \circ (f~p_4) \circ (f~p_3) \circ (f~p_2) \circ (\lambda v. f~v~x))\hskip1mm p_1\\ {\Delta}p_2 = - \eta\hskip1mm D(C(g~x) \circ (f~p_4) \circ (f~p_3) \circ (\lambda v. f~v(f~p_1~x)))\hskip1mm p_2\\ {\Delta}p_3 = - \eta\hskip1mm D(C(g~x) \circ (f~p_4) \circ (\lambda v. f~v(f~p_2(f~p_1~x))))\hskip1mm p_3\\ {\Delta}p_4 = - \eta\hskip1mm D(C(g~x) \circ (\lambda v. f~v(f~p_3(f~p_2(f~p_1~x)))))\hskip1mm p_4\\ \end{array}\)
This shows that to calculate \({\Delta}p_i\) one must calculate \(D\) applied to the composition of functions. This is done with the chain rule for calculating the derivatives of compositions of functions.
The chain rule states that for arbitrary differentiable functions \(\theta_1 : {\Bbb R}\rightarrow{\Bbb R}\) and \(\theta_2 : {\Bbb R}\rightarrow{\Bbb R}\):
\(D(\theta_1 \circ \theta_2) = (D \theta_1 \circ \theta_2) \cdot D \theta_2\)
Where “\(\cdot\)” is pointwise multiplication of functions. Thus if “\(\times\)” is the usual multiplication on real numbers, then applying both sides of this equation to \(x\) gives
\(\begin{array}{l} D(\theta_1\circ\theta_2)x\\ ~ = ((D \theta_1\circ\theta_2) \cdot D \theta_2)x\\ ~ = ((D \theta_1\circ\theta_2)x) {\times} (D \theta_2~x)\\ ~ = (D \theta_1(\theta_2~x)) {\times} (D \theta_2~x)\\ ~ = D \theta_1(\theta_2~x) {\times} D \theta_2~x \end{array}\)
The example below shows the rule works in a specific case and might help build an intuition for why it works in general. Let \(\theta_1\) and \(\theta_2\) be defined by
\(\begin{array}{l} \theta_1(x) = x^m\\ \theta_2(x) = x^n\\ \end{array}\)
then by the rule for differentiating powers:
\(\begin{array}{l} D\theta_1(x) = m x^{m-1}\\ D\theta_2(x) = n x^{n-1}\\ \end{array}\)
By the definition of function composition
\(\begin{array}{l} (\theta_1 \circ \theta_2)(x) = \theta_1(\theta_2~x) = (\theta_2~x)^m = (x^n)^m = x^{(mn)} \end{array}\)
so by the rule for differentiating powers: \(D(\theta_1 \circ \theta2)(x) = (mn) x^{(mn) -1}\).
Also
\(\begin{array}{l} D \theta_1(\theta_2~x) {\times} D \theta_2(x) = m (\theta_2~x)^{m-1} {\times} n x^{n-1}\\ \phantom{D \theta_1(\theta_2~x) {\times} D \theta_2(x)} = m (x^n)^{m-1} {\times} n x^{n-1}\\ \phantom{D \theta_1(\theta_2~x) {\times} D \theta_2(x)} = m x^{n{\times}(m-1)} {\times} n x^{n-1}\\ \phantom{D \theta_1(\theta_2~x) {\times} D \theta_2(x)} = mn x^{n{\times}(m-1) + (n-1)}\\ \phantom{D \theta_1(\theta_2~x) {\times} D \theta_2(x)} = mn x^{n{\times}m - n{\times}1 + n - 1}\\ \phantom{D \theta_1(\theta_2~x) {\times} D \theta_2(x)} = mn x^{(nm) - 1}\\ \end{array}\)
so for this example \(D(\theta_1 \circ \theta_2)(x) = D \theta_1(\theta_2~x) {\times} D \theta_2(x)\). For the general case, see the Wikipedia article, which has three proofs of the chain rule.
The equation \(D(\theta_1 \circ \theta_2)(x) = D \theta_1(\theta_2~x) {\times} D \theta_2(x)\) can be used recursively, after left-associating multiple function compositions, for example:
\(\begin{array}{l} D(\theta_1 \circ \theta_2 \circ \theta_3)~x = D((\theta_1 \circ \theta_2) \circ \theta_3)~x \\ \phantom{D(\theta_1 \circ \theta_2 \circ \theta_3)~x} = D(\theta_1 \circ \theta_2) (\theta_3~x) \times D \theta_3~x\\ \phantom{D(\theta_1 \circ \theta_2 \circ \theta_3)~x} = D\theta_1(\theta_2(\theta_3~x)) \times D\theta_2(\theta_3~x) \times D \theta_3~x\\ \end{array}\)
Consider now \(y = E_1(E_2)\), where \(E_2\) is an expression involving \(x\). By introducing a new variable, \(u\) say, this can be split into two equations \(y = E_1(u)\) and \(u=E_2\).
As an example consider \(y= sin(x^2)\). This can be split into the two equations \(y=\sin(u)\) and \(u=x^2\).
The derivative equations are then \(\frac{dy}{du} = \cos(u)\) and \(\frac{du}{dx} = 2x\).
The chain rule in Leibitz notation is \(\frac{dy}{dx} = \frac{dy}{du} \times \frac{du}{dx}\).
Applying this to the example gives \(\frac{dy}{dx} = \cos(u) \times 2x\).
As \(u=x^2\) this is \(\frac{dy}{dx} = \cos(x^2) \times 2x\).
To compare this calculation with the equivalent one using Euler notation, let \(\phi_1 = \lambda x E_1\) and \(\phi_2 = \lambda x E_2\). Then \(D(\lambda x. E_1(E_2)) = D(\phi_1 \circ \phi_2)\) and so \(D(\lambda x. E_1(E_2)) = (D\phi_1 \circ \phi_2) \cdot D\phi_2\) by the chain rule. The Euler notation calculation for the example is:
\(\begin{array}{ll} D(\lambda x. E_1(E_2))x &=~~~ ((D\phi_1 \circ \phi_2) \cdot D\phi_2)~x \\ &=~~~ ((D\phi_1 \circ \phi_2) \cdot D\phi_2)~x \\ &=~~~ (D\phi_1 \circ \phi_2)~x \times D\phi_2~x \\ &=~~~ D\phi_1(\phi_2~x) \times D\phi_2~x \\ &=~~~ D \sin(x^2) \times D(\lambda x. x^2)~x \\ &=~~~ \cos(x^2) \times (\lambda x. 2 x)~x \\ &=~~~ \cos(x^2) \times 2 x \end{array}\)
I think it’s illuminating – at least for someone like me whose calculus is very rusty – to compare the calculation of the deltas for the four-neuron linear network using both Euler and Leibniz notation. This is done in the next two sections.
Note. Starting here, and continuing for the rest of this article, I give a lot of detailed and sometimes repetitive calculations. The aim is to make regularities explicit so that the transition to more compact notations where the regularities are implicit – e.g. using vectors and matrices – is easy to explain.
First, the activations \(a_1\), \(a_2\), \(a_3\), \(a_4\) are calculated by making a forward pass through the network.
\(\begin{array}{l} a_1 = f~p_1~x \\ a_2 = f~p_2~a_1 \\ a_3 = f~p_3~a_2 \\ a_4 = f~p_4~a_3 \\ \end{array}\)
Plugging \(a_1\), \(a_2\), \(a_3\) into the equation for each the \({\Delta}p_i\) results in:
\(\begin{array}{l} {\Delta}p_1 = - \eta\hskip1mm D(C(g~x) \circ (f~p_4) \circ (f~p_3) \circ (f~p_2) \circ (\lambda v. f~v~x))\hskip1mm p_1\\ {\Delta}p_2 = - \eta\hskip1mm D(C(g~x) \circ (f~p_4) \circ (f~p_3) \circ (\lambda v. f~v~a_1))\hskip1mm p_2\\ {\Delta}p_3 = - \eta\hskip1mm D(C(g~x) \circ (f~p_4) \circ (\lambda v. f~v~a_2))\hskip1mm p_3\\ {\Delta}p_4 = - \eta\hskip1mm D(C(g~x) \circ (\lambda v. f~v~a_3))\hskip1mm p_4\\ \end{array}\)
Working backwards, these equation are then recursively evaluated using the chain rule. With \(i\) counting down from 4 to 1, compute \({{\hskip0.5mm\mathsf{d}}}_i\) and \({\Delta}p_i\) as follows.
\({{\hskip0.5mm\mathsf{d}}}_4 = D(C(g{\hskip0.8mm}x))a_4\)
\(\begin{array}{l} {\Delta}p_4 = {-}\eta{\times} D(C(g{\hskip0.8mm}x) \circ (\lambda v.f{\hskip0.8mm}v{\hskip0.8mm}a_3))p_4\\ \phantom{{\Delta}p_4} = {-}\eta{\times} D(C(g{\hskip0.8mm}x)) (f{\hskip0.8mm}p_4{\hskip0.8mm}a_3) {\times} D(\lambda v.f{\hskip0.8mm}v{\hskip0.8mm}a_3)p_4\\ \phantom{{\Delta}p_4} = {-}\eta{\times} D(C(g{\hskip0.8mm}x)) a_4 {\times} D(\lambda v.f{\hskip0.8mm}v{\hskip0.8mm}a_3)p_4\\ \phantom{{\Delta}p_4} = {-}\eta{\times} {{\hskip0.5mm\mathsf{d}}}_4 {\times} D(\lambda v.f{\hskip0.8mm}v{\hskip0.8mm}a_3)p_4\\ \end{array}\)
\({{\hskip0.5mm\mathsf{d}}}_3 = D(C(g{\hskip0.8mm}x))a_4 {\times} D(f{\hskip0.8mm}p_4)a_3 = {{\hskip0.5mm\mathsf{d}}}_4 {\times} D(f{\hskip0.8mm}p_4)a_3\)
\(\begin{array}{l} {\Delta}p_3 = {-}\eta{\times} D(C(g{\hskip0.8mm}x) \circ (f{\hskip0.8mm}p_4) \circ (\lambda v.f{\hskip0.8mm}v{\hskip0.8mm}a_2))p_3\\ \phantom{{\Delta}p_3} = {-}\eta{\times} D(C(g{\hskip0.8mm}x) \circ (f{\hskip0.8mm}p_4))(f{\hskip0.8mm}p_3{\hskip0.8mm}a_2) {\times} D(\lambda v.f{\hskip0.8mm}v{\hskip0.8mm}a_2)p_3\\ \phantom{{\Delta}p_3} = {-}\eta{\times} D(C(g{\hskip0.8mm}x) \circ (f{\hskip0.8mm}p_4))a_3 {\times} D(\lambda v.f{\hskip0.8mm}v{\hskip0.8mm}a_2)p_3\\ \phantom{{\Delta}p_3} = {-}\eta{\times} D(C(g{\hskip0.8mm}x)) ((f{\hskip0.8mm}p_4)a_3) {\times} D(f{\hskip0.8mm}p_4)a_3 {\times} D(\lambda v.f{\hskip0.8mm}v{\hskip0.8mm}a_2)p_3\\ \phantom{{\Delta}p_3} = {-}\eta{\times} D(C(g{\hskip0.8mm}x)) a_4 {\times} D(f{\hskip0.8mm}p_4)a_3 {\times} D(\lambda v.f{\hskip0.8mm}v{\hskip0.8mm}a_2)p_3\\ \phantom{{\Delta}p_3} = {-}\eta{\times} {{\hskip0.5mm\mathsf{d}}}_4 {\times} D(f{\hskip0.8mm}p_4)a_3 {\times} D(\lambda v.f{\hskip0.8mm}v{\hskip0.8mm}a_2)p_3\\ \phantom{{\Delta}p_3} = {-}\eta{\times} {{\hskip0.5mm\mathsf{d}}}_3 {\times} D(\lambda v.f{\hskip0.8mm}v{\hskip0.8mm}a_2)p_3\\ \end{array}\)
\({{\hskip0.5mm\mathsf{d}}}_2 = D(C(g{\hskip0.8mm}x))a_4 {\times} D(f{\hskip0.8mm}p_4)a_3 {\times} D(f{\hskip0.8mm}p_3)a_2 = {{\hskip0.5mm\mathsf{d}}}_3 {\times} D(f{\hskip0.8mm}p_3)a_2\)
\(\begin{array}{l} {\Delta}p_2 = {-}\eta{\times} D(C(g{\hskip0.8mm}x) \circ (f{\hskip0.8mm}p_4) \circ (f{\hskip0.8mm}p_3) \circ (\lambda v.f{\hskip0.8mm}v{\hskip0.8mm}a_1))p_2\\ \phantom{{\Delta}p_2} = {-}\eta{\times} D(C(g{\hskip0.8mm}x) \circ (f{\hskip0.8mm}p_4) \circ (f{\hskip0.8mm}p_3))(f{\hskip0.8mm}p_2{\hskip0.8mm}a_1) {\times} D(\lambda v.f{\hskip0.8mm}v{\hskip0.8mm}a_1)p_2\\ \phantom{{\Delta}p_2} = {-}\eta{\times} D(C(g{\hskip0.8mm}x) \circ (f{\hskip0.8mm}p_4) \circ (f{\hskip0.8mm}p_3))a_2 {\times} D(\lambda v.f{\hskip0.8mm}v{\hskip0.8mm}a_1)p_2\\ \phantom{{\Delta}p_2} = {-}\eta{\times} D(C(g{\hskip0.8mm}x) \circ (f{\hskip0.8mm}p_4)) ((f{\hskip0.8mm}p_3)a_2) {\times} D(f{\hskip0.8mm}p_3)a_2 {\times} D(\lambda v.f{\hskip0.8mm}v{\hskip0.8mm}a_1)p_2\\ \phantom{{\Delta}p_2} = {-}\eta{\times} D(C(g{\hskip0.8mm}x) \circ (f{\hskip0.8mm}p_4)) a_3 {\times} D(f{\hskip0.8mm}p_3)a_2 {\times} D(\lambda v.f{\hskip0.8mm}v{\hskip0.8mm}a_1)p_2\\ \phantom{{\Delta}p_2} = {-}\eta{\times} D(C(g{\hskip0.8mm}x)) ((f{\hskip0.8mm}p_4)a_3) {\times} D(f{\hskip0.8mm}p_4)a_3 {\times} D(f{\hskip0.8mm}p_3)a_2 {\times} D(\lambda v.f{\hskip0.8mm}v{\hskip0.8mm}a_1)p_2\\ \phantom{{\Delta}p_2} = {-}\eta{\times} {{\hskip0.5mm\mathsf{d}}}_2 {\times} D(\lambda v.f{\hskip0.8mm}v{\hskip0.8mm}a_1)p_2\\ \end{array}\)
\({{\hskip0.5mm\mathsf{d}}}_1 = D(C(g{\hskip0.8mm}x))a_4 {\times} D(f{\hskip0.8mm}p_4)a_3 {\times} D(f{\hskip0.8mm}p_3)a_2 {\times} D(f{\hskip0.8mm}p_2)a_1 = {{\hskip0.5mm\mathsf{d}}}_2 {\times} D(f{\hskip0.8mm}p_2)a_1\)
\(\begin{array}{l} {\Delta}p_1 = {-}\eta{\times} D(C(g{\hskip0.8mm}x) \circ (f{\hskip0.8mm}p_4) \circ (f{\hskip0.8mm}p_3) \circ (f{\hskip0.8mm}p_2) \circ (\lambda v.f{\hskip0.8mm}v{\hskip0.8mm}x))p_1\\ \phantom{{\Delta}p_1} = {-}\eta{\times} D((C(g{\hskip0.8mm}x) \circ (f{\hskip0.8mm}p_4) \circ (f{\hskip0.8mm}p_3) \circ (f{\hskip0.8mm}p_2)) \circ (\lambda v.f{\hskip0.8mm}v{\hskip0.8mm}x))p_1\\ \phantom{{\Delta}p_1} = {-}\eta{\times} D(C(g{\hskip0.8mm}x) \circ (f{\hskip0.8mm}p_4) \circ (f{\hskip0.8mm}p_3) \circ (f{\hskip0.8mm}p_2)) (f{\hskip0.8mm}p_1{\hskip0.8mm}x) {\times} D(\lambda v.f{\hskip0.8mm}v{\hskip0.8mm}x)p_1\\ \phantom{{\Delta}p_1} = {-}\eta{\times} D((C(g{\hskip0.8mm}x) \circ (f{\hskip0.8mm}p_4) \circ (f{\hskip0.8mm}p_3)) \circ (f{\hskip0.8mm}p_2)) a_1 {\times} D(\lambda v.f{\hskip0.8mm}v{\hskip0.8mm}x)p_1\\ \phantom{{\Delta}p_1} = {-}\eta{\times} D(C(g{\hskip0.8mm}x) \circ (f{\hskip0.8mm}p_4) \circ (f{\hskip0.8mm}p_3)) (f{\hskip0.8mm}p_2{\hskip0.8mm}a_1) {\times} D(f{\hskip0.8mm}p_2) a_1 {\times} D(\lambda v.f{\hskip0.8mm}v{\hskip0.8mm}x)p_1\\ \phantom{{\Delta}p_1} = {-}\eta{\times} D(C(g{\hskip0.8mm}x) \circ (f{\hskip0.8mm}p_4) \circ (f{\hskip0.8mm}p_3)) a_2 {\times} D(f{\hskip0.8mm}p_2)a_1 {\times} D(\lambda v.f{\hskip0.8mm}v{\hskip0.8mm}x)p_1\\ \phantom{{\Delta}p_1} = {-}\eta{\times} D(C(g{\hskip0.8mm}x) \circ (f{\hskip0.8mm}p_4)) (f{\hskip0.8mm}p_3{\hskip0.8mm}a_2) {\times} D(f{\hskip0.8mm}p_3) a_2 {\times} D(f{\hskip0.8mm}p_2)a_1 {\times} D(\lambda v.f{\hskip0.8mm}v{\hskip0.8mm}x)p_1\\ \phantom{{\Delta}p_1} = {-}\eta{\times} D(C(g{\hskip0.8mm}x) \circ (f{\hskip0.8mm}p_4)) a_3 {\times} D(f{\hskip0.8mm}p_3)a_2 {\times} D(f{\hskip0.8mm}p_2)a_1 {\times} D(\lambda v.f{\hskip0.8mm}v{\hskip0.8mm}x)p_1\\ \phantom{{\Delta}p_1} = {-}\eta{\times} D(C(g{\hskip0.8mm}x)) (f{\hskip0.8mm}p_4{\hskip0.8mm}a_3) {\times} D(f{\hskip0.8mm}p_4) a_3 {\times} D(f{\hskip0.8mm}p_3)a_2 {\times} D(f{\hskip0.8mm}p_2)a_1 {\times} D(\lambda v.f{\hskip0.8mm}v{\hskip0.8mm}x)p_1\\ \phantom{{\Delta}p_1} = {-}\eta{\times} D(C(g{\hskip0.8mm}x)) a_4 {\times} D(f{\hskip0.8mm}p_4)a_3 {\times} D(f{\hskip0.8mm}p_3)a_2 {\times} D(f{\hskip0.8mm}p_2)a_1 {\times} D(\lambda v.f{\hskip0.8mm}v{\hskip0.8mm}x)p_1\\ \phantom{{\Delta}p_1} = {-}\eta{\times} {{\hskip0.5mm\mathsf{d}}}_1 {\times} D(\lambda v.f{\hskip0.8mm}v{\hskip0.8mm}x)p_1\\ \end{array}\)
Summarising the results of the calculation using Euler notation and omitting “\(\times\)”:
\(\begin{array}{ll} {{\hskip0.5mm\mathsf{d}}}_4 = D(C(g{\hskip0.8mm}x))a_4 \qquad\qquad & {\Delta}p_4 = -\eta {{\hskip0.5mm\mathsf{d}}}_4 D(\lambda v.f{\hskip1mm}v{\hskip1mm}a_3) p_4\\ {{\hskip0.5mm\mathsf{d}}}_3 = {{\hskip0.5mm\mathsf{d}}}_4 { } D(f{\hskip0.8mm}p_4)a_3 & {\Delta}p_3 = -\eta {{\hskip0.5mm\mathsf{d}}}_3 D(\lambda v.f{\hskip1mm}v{\hskip1mm}a_2) p_3\\ {{\hskip0.5mm\mathsf{d}}}_2 = {{\hskip0.5mm\mathsf{d}}}_3 { } D(f{\hskip0.8mm}p_3)a_2 & {\Delta}p_2 = -\eta {{\hskip0.5mm\mathsf{d}}}_2 D(\lambda v.f{\hskip1mm}v{\hskip1mm}a_1) p_2\\ {{\hskip0.5mm\mathsf{d}}}_1 = {{\hskip0.5mm\mathsf{d}}}_2 { } D(f{\hskip0.8mm}p_2)a_1 & {\Delta}p_1 = -\eta {{\hskip0.5mm\mathsf{d}}}_1 D(\lambda v.f{\hskip1mm}v{\hskip1mm}x) p_1\\ \end{array}\)
The equations for the activations are
\(~~~\begin{array}{l} a_1 = f~p_1~x \\ a_2 = f~p_2~a_1 \\ a_3 = f~p_3~a_2 \\ a_4 = f~p_4~a_3 \\ \end{array}\)
These specify equations between variables \(a_1\), \(a_2\), \(a_3\), \(a_4\), \(p_1\), \(p_2\), \(p_3\), \(p_4\) (\(x\) is considered a constant).
Let \(c\) be a variable representing the cost.
\(c = \phi(p_1,p_2,p_3,p_4) = C(g~x)(f~p_4(f~p_3(f~p_2(f~p_1~x)))) = C (g~x) a_4\)
In leibniz notation \(\nabla\phi(p_1,p_2,p_3,p_4) = (\frac{\partial c}{\partial p_1}, \frac{\partial c}{\partial p_2}, \frac{\partial c}{\partial p_3}, \frac{\partial c}{\partial p_4})\) and so \({\Delta}p_i = -\eta\hskip1mm \frac{\partial c}{\partial p_i}\).
By the chain rule:
\(\frac{\partial c}{\partial p_1} = \frac{\partial c}{\partial a_4} \frac{\partial a_4}{\partial a_3} \frac{\partial a_3}{\partial a_2} \frac{\partial a_2}{\partial a_1} \frac{\partial a_1}{\partial p_1}\)
\(\frac{\partial c}{\partial p_2} = \frac{\partial c}{\partial a_4} \frac{\partial a_4}{\partial a_3} \frac{\partial a_3}{\partial a_2} \frac{\partial a_2}{\partial p_2}\)
\(\frac{\partial c}{\partial p_3} = \frac{\partial c}{\partial a_4} \frac{\partial a_4}{\partial a_3} \frac{\partial a_3}{\partial p_3}\)
\(\frac{\partial c}{\partial p_4} = \frac{\partial c}{\partial a_4} \frac{\partial a_4}{\partial p_4}\)
Calculate as follows:
\({{\hskip0.5mm\mathsf{d}}}_4 = \frac{\partial c}{\partial a_4} = D(C(g~x))a_4\)
\({\Delta}p_4 = - \eta\hskip1mm \frac{\partial c}{\partial a_4} \frac{\partial a_4}{\partial p_4}\\ \phantom{{\Delta}p_4} = - \eta\hskip1mm {{\hskip0.5mm\mathsf{d}}}_4 D(\lambda v. f~v~a_3) p_4\)
\({{\hskip0.5mm\mathsf{d}}}_3 = \frac{\partial c}{\partial a_4} \frac{\partial a_4}{\partial a_3} = {{\hskip0.5mm\mathsf{d}}}_4 D(f~p_3)a_3\)
\({\Delta}p_3 = - \eta\hskip1mm \frac{\partial c}{\partial a_4} \frac{\partial a_4}{\partial a_3} \frac{\partial a_3}{\partial p_3}\\ \phantom{{\Delta}p_3} =- \eta\hskip1mm {{\hskip0.5mm\mathsf{d}}}_3 D(\lambda v. f~v~a_2)p_3\)
\({{\hskip0.5mm\mathsf{d}}}_2 = \frac{\partial c}{\partial a_4} \frac{\partial a_4}{\partial a_3} \frac{\partial a_3}{\partial a_2} = {{\hskip0.5mm\mathsf{d}}}_3 D(f~p_2)a_2\)
\({\Delta}p_2 = - \eta\hskip1mm \frac{\partial c}{\partial a_4} \frac{\partial a_4}{\partial a_3} \frac{\partial a_3}{\partial a_2} \frac{\partial a_2}{\partial p_2}\\ \phantom{{\Delta}p_2} = - \eta\hskip1mm {{\hskip0.5mm\mathsf{d}}}_2 D(f~a_1)p_2\)
\({{\hskip0.5mm\mathsf{d}}}_1 = \frac{\partial c}{\partial a_4} \frac{\partial a_4}{\partial a_3} \frac{\partial a_3}{\partial a_2} \frac{\partial a_2}{\partial a_1} = {{\hskip0.5mm\mathsf{d}}}_2 D(f~p_2)a_1\)
\({\Delta}p_1 = - \eta\hskip1mm \frac{\partial c}{\partial a_4} \frac{\partial a_4}{\partial a_3} \frac{\partial a_3}{\partial a_2} \frac{\partial a_2}{\partial a_1} \frac{\partial a_1}{\partial p_1}\\ \phantom{{\Delta}p_1} = - \eta\hskip1mm {{\hskip0.5mm\mathsf{d}}}_1 D(\lambda v. f~v~x)p_1\)
Summarising the results of the calculation using Leibniz notation:
\(\begin{array}{ll} {{\hskip0.5mm\mathsf{d}}}_4 = D(C(g~x)) \qquad\qquad & {\Delta}p_4 = - \eta\hskip1mm {{\hskip0.5mm\mathsf{d}}}_4 D(\lambda v. f~v~a_3) p_4\\ {{\hskip0.5mm\mathsf{d}}}_3 = {{\hskip0.5mm\mathsf{d}}}_4 D(f~p_3)a_3 & {\Delta}p_3 =- \eta\hskip1mm {{\hskip0.5mm\mathsf{d}}}_3 D(\lambda v. f~v~a_2)p_3\\ {{\hskip0.5mm\mathsf{d}}}_2 = {{\hskip0.5mm\mathsf{d}}}_3 D(f~p_2)a_2 & {\Delta}p_2 = - \eta\hskip1mm {{\hskip0.5mm\mathsf{d}}}_2 D(\lambda v. f~v~a_1)p_2\\ {{\hskip0.5mm\mathsf{d}}}_1 = {{\hskip0.5mm\mathsf{d}}}_2 D(f~p_2)a_1 & {\Delta}p_1 = - \eta\hskip1mm {{\hskip0.5mm\mathsf{d}}}_1 D(\lambda v. f~v~x)p_1 \end{array}\)
Fortunately, the calculations using Euler and Leibniz notations gave the same results. From these the following two-pass pseudocode algorithm can be used to calculate the deltas.
\(~~~\begin{array}{ll} a_1 &=~~~ f~p_1~x\texttt{;} \\ a_2 &=~~~ f~p_2~a_1\texttt{;} \\ a_3 &=~~~ f~p_3~a_2\texttt{;} \\ a_4 &=~~~ f~p_4~a_3\texttt{;} \\ \end{array}\)
\(~~~\begin{array}{ll} {{\hskip0.5mm\mathsf{d}}}_4 &=~~~ D(C(g~x))a_4\texttt{;} \\ {\Delta}p_4 &=~~~ -\eta {{\hskip0.5mm\mathsf{d}}}_4 D(\lambda v.f~v~a_3) p_4\texttt{;}\\ {{\hskip0.5mm\mathsf{d}}}_3 &=~~~ {{\hskip0.5mm\mathsf{d}}}_4 D(f~p_4)a_3\texttt{;} \\ {\Delta}p_3 &=~~~ -\eta {{\hskip0.5mm\mathsf{d}}}_3 D(\lambda v.f~v~a_2) p_3\texttt{;}\\ {{\hskip0.5mm\mathsf{d}}}_2 &=~~~ {{\hskip0.5mm\mathsf{d}}}_3 D(f~p_3)a_2\texttt{;} \\ {\Delta}p_2 &=~~~ -\eta {{\hskip0.5mm\mathsf{d}}}_2 D(\lambda v.f~v~a_1) p_2\texttt{;}\\ {{\hskip0.5mm\mathsf{d}}}_1 &=~~~ {{\hskip0.5mm\mathsf{d}}}_2 D(f~p_2)a_1\texttt{;} \\ {\Delta}p_1 &=~~~ -\eta {{\hskip0.5mm\mathsf{d}}}_1 D(\lambda v.f~v~x) p_1\texttt{;} \\ \end{array}\)
So far the cost function \(C\) which measures the error of a network output, and the function \(f\) modelling the behaviour of neurons, have not been specified. The next section discusses the actual functions these are.
Here is an example of a typical neuron taken from Chapter 1 of Nielsen’s book.
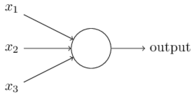
This example has three inputs, but neurons may have any number of inputs. A simple model of the behaviour of this three-input neuron is given by the equation
\(\begin{array}{ll} \mathrm{output} & \hskip-3mm = \sigma(w_1 x_1 + w_2 x_2 + w_3 x_3 + b) \end{array}\)
where \(w_1\), \(w_2\) and \(w_3\) are weights and \(b\) is the bias. These are real numbers and are the parameters of the neuron that machine learning aims to learn. The number of weights equals the number of inputs, so in the linear network example above each neuron only had one parameter (the bias was ignored).
The function \(\sigma\) is a sigmoid function, i.e. has a graph with the shape shown below.
There are explanations of why sigmoid functions are used in Chapter 1 of Nielsen’s book and Patrick Winston’s online course.
The choice of a particular sigmoid function \(\sigma\) – called an activation function – depends on the application. One possibility is the trigonometric \(\tanh\) function; a more common choice, at least in elementary expositions of neural networks, is the logistic function, perhaps because it has nice mathematical properties like a simple derivative. The logistic function is defined by:
\(\sigma(x)=\frac {1}{1+e^{-x}}\)
The derivative \(D \sigma\) or \(\sigma'\) is discussed in the section entitled Derivative of the activation function below.
The graph of \(\sigma\) below is taken from the Wikipedia article on the logistic function.
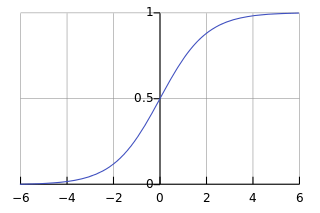
I also found online the graph of \(\tanh\) below.
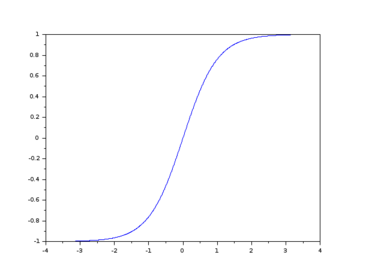
Both the logistic and \(\tanh\) functions have a sigmoid shape and they are also related by the equation \(\tanh(x)= 2\hskip1mm \sigma(2\hskip1mm x)-1\). The derivative of \(\tanh\) is pretty simple, so I don’t really know how one chooses between \(\sigma\) and \(\tanh\), but Google found a discussion on Quora on the topic.
The logistic function \(\sigma\) is used here, so the behaviour of the example neuron is
\(\begin{array}{ll} \mathrm{output} & \hskip-3mm = \sigma(w_1 x_1 + w_2 x_2 + w_3 x_3 + b)\\ & \hskip-3mm = \frac{1}{1+e^{-(w_1 x_1 + w_2 x_2 + w_3 x_3 + b)}}\\ & \hskip-3mm = \frac{1}{1+exp(-(w_1 x_1 + w_2 x_2 + w_3 x_3 + b))} \end{array}\)
where the last line is just the one above it rewritten using the notation \(exp(v)\) instead of \(e^v\) for exponentiation.
The other uninterpreted function used above is the cost function \(C\). This measures the closeness of a desired output \(g~x\) for an input \(x\) to the output actually produced by the network, i.e. \(f_{\scriptsize\textsf{N}}~p~x\).
Various particular cost functions are used, but the usual choice in elementary expositions is the mean squared error (MSE) defined by \(C~u~v = \frac{1}{2}(u-v)^2\), so the error for a given input \(x\) is
\(C(g~x)(f_{\scriptsize\textsf{N}}~p~x) = \frac{1}{2} (g~x - f_{\scriptsize\textsf{N}}~p~x)^2\)
Note that \(C\) is commutative: \(C~u~v = C~v~u\). However, although, for example \(C~2~3 = C~3~2\), the partially applied functions \(C~2\) and \(C~3\) are not equal.
Using logistic neuron models, the behaviour each neuron in the four-neuron linear network is given by the function \(f\) defined by
\(f~w~a = \frac{1}{1+exp(-(w a + b))}\)
where the weight \(w\) is the only parameter and (temporarily) the bias \(b\) is not being considered as a parameter to be learnt, but is treated as a constant.
Using mean square error to measure cost, the function to be minimised is
\(\phi(w_1,w_2,w_3,w_4) = \frac{1}{2} (f~w_4 (f~w_3 (f~w_2 (f~w_1~x))) - g~x)^2\)
For an input \(x\), the forward pass computation of the activations \(a_i\) is
\(~~~\begin{array}{ll} a_1 &=~~~ f~w_1~x\texttt{;} \\ a_2 &=~~~ f~w_2~a_1\texttt{;} \\ a_3 &=~~~ f~w_3~a_2\texttt{;} \\ a_4 &=~~~ f~w_4~a_3\texttt{;} \\ \end{array}\)
Expanding the definition of \(f\):
\(\begin{array}{l} a_1 = \sigma(w_1 x + b) ~= \frac{1}{1+exp(-(w_1 x + b))}\\ a_2 = \sigma(w_2 a_1 + b) = \frac{1}{1+exp(-(w_2 a_1 + b))}\\ a_3 = \sigma(w_3 a_2 + b) = \frac{1}{1+exp(-(w_3 a_2 + b))}\\ a_4 = \sigma(w_4 a_3 + b) = \frac{1}{1+exp(-(w_4 a_3 + b))}\\ \end{array}\)
In the instantiated diagram below the weights are shown on the incoming arrows and the bias inside the nodes (the logistic function \(\sigma\) is assumed for the sigmoid function).
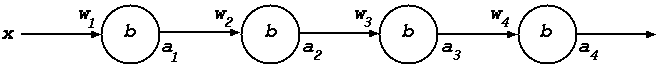
To execute the backward pass one must compute derivatives, specifically: \(D(C(G~x))\) and \(D(\lambda v.f~v~x)\) for the input \(x\), \(D(\lambda v.f~v~a_i)\) for \(i=1,2,3\) and \(D(f~w_i)\) for \(i=2,3,4\).
The particular functions whose derivatives are needed are:
\(\begin{array}{ll} \lambda v. \frac{1}{2}(v - g~x)^2 & (x \textrm{ is treated as a constant and } g \textrm{ as a constant function})\\ \lambda v. \frac{1}{1+exp(-(v a + b))} & (a \textrm{ and } b \textrm{ are treated as constants})\\ \lambda v. \frac{1}{1+exp(-(w v + b))} & (w \textrm{ and } b \textrm{ are treated as constants})\\ \end{array}\)
Note that the second and third functions above are essentially the same as multiplication is commutative.
The cost function is \(\lambda v. \frac{1}{2}(u - v)^2\). The derivative will be calculated using Leibniz notation, starting with the equation.
\(c = \frac{1}{2}(u - v)^2\)
Split this into three equations using two new variables, \({\alpha}\) and \({\beta}\), and then differentiate each equation using the standard rules - see table below.
\(\begin{array}{l|l|l} \hline {\textbf{Equation}} & {\textbf{Derivative}} & {\textbf{Differentiation rules used}} \\ \hline c = \frac{1}{2}{\alpha} & \frac{dc}{d{\alpha}} = \frac{1}{2} & \textrm{multiplication by constant rule, } \frac{d{\alpha}}{d{\alpha}} = 1\\ \hline {\alpha} = {\beta}^2 & \frac{d{\alpha}}{d{\beta}} = 2 {\beta} & \textrm{power rule}\\ \hline {\beta} = u - v & \frac{d{\beta}}{dv} = -1 & \textrm{subtraction rule}, \frac{du}{dv}{=} 0, \frac{dv}{dv}{=}1\\ \hline \end{array}\)
Hence \(\frac{dc}{dv} = \frac{dc}{d{\alpha}} \times \frac{d{\alpha}}{d{\beta}} \times \frac{d{\beta}}{dv} = \frac{1}{2} \times 2 {\beta} \times -1 = \frac{1}{2} \times 2 (u - v) \times -1 = v - u\)
Thus \(D(\lambda v. \frac{1}{2}(u - v)^2) = \lambda v. v - u\).
The other function whose derivative is needed is \(\lambda v. \sigma(w v + b) = \lambda v. \frac{1}{1+exp(-(w v + b))}\).
This is \(\lambda v. \sigma(w v + b)\), where \(\sigma\) is the logistic function. \(D \sigma = \lambda v. \sigma(v)(1 - \sigma(v))\). The easy derivation of the derivative of the logistic function \(\sigma\) can be found here. Using Lagrange notation \(D\sigma\) is \(\sigma'\).
The derivative will be calculated using Leibniz notation, starting with the equation
\(a = \sigma(w v + b)\)
where \(a\) is dependent on \(v\). Split this into two equations using a new variables \({z}\) and then differentiate each equation - see table below.
\(\begin{array}{l|l|l} \hline {\textbf{Equation}} & {\textbf{Derivative}} & {\textbf{Differentiation rules used}} \\ \hline a = \sigma({z}) & \frac{da}{d{z}} = \sigma({z})(1{-}\sigma({z})) & \textrm{see discussion above} \\ \hline {z} = w v + b & \frac{d{z}}{dv} = w & \textrm{addition, constant multiplication}, \frac{dv}{dv}{=}1, \frac{db}{dv}{=}0\\ \hline \end{array}\)
Hence \(\frac{da}{dv} = \frac{da}{d{z}} \times \frac{d{z}}{dv} = \sigma({z})(1{-}\sigma({z})) \times w = \sigma(w v + b)(1{-}\sigma(w v + b)) \times w\)
Thus \(D(\lambda v. \sigma(w v + b)) = \lambda v. \sigma(w v + b)(1 - \sigma(w v + b)) \times w\).
Since \(D \sigma = \lambda v. \sigma(v)(1 - \sigma(v))\), the equation above can be written more compactly as \(D(\lambda v. \sigma(w v + b)) = \lambda v. D\sigma(w v + b) \times w\) or even more compactly by writing \(\sigma'\) for \(D\sigma\) and omitting the “\(\times\)” symbol: \(D(\lambda v. \sigma(w v + b)) = \lambda v. \sigma'(w v + b) w\).
Recall the pseudocode algorithm given earlier.
Forward pass:
\(~~~\begin{array}{ll} a_1 &=~~~ f~p_1~x\texttt{;} \\ a_2 &=~~~ f~p_2~a_1\texttt{;} \\ a_3 &=~~~ f~p_3~a_2\texttt{;} \\ a_4 &=~~~ f~p_4~a_3\texttt{;} \\ \end{array}\)
Backward pass:
\(~~~\begin{array}{ll} {{\hskip0.5mm\mathsf{d}}}_4 &=~~~ D(C(g~x))a_4\texttt{;} \\ {\Delta}p_4 &=~~~ -\eta {{\hskip0.5mm\mathsf{d}}}_4 D(\lambda v.f~v~a_3) p_4\texttt{;}\\ {{\hskip0.5mm\mathsf{d}}}_3 &=~~~ {{\hskip0.5mm\mathsf{d}}}_4 D(f~p_4)a_3\texttt{;} \\ {\Delta}p_3 &=~~~ -\eta {{\hskip0.5mm\mathsf{d}}}_3 D(\lambda v.f~v~a_2) p_3\texttt{;}\\ {{\hskip0.5mm\mathsf{d}}}_2 &=~~~ {{\hskip0.5mm\mathsf{d}}}_3 D(f~p_3)a_2\texttt{;} \\ {\Delta}p_2 &=~~~ -\eta {{\hskip0.5mm\mathsf{d}}}_2 D(\lambda v.f~v~a_1) p_2\texttt{;}\\ {{\hskip0.5mm\mathsf{d}}}_1 &=~~~ {{\hskip0.5mm\mathsf{d}}}_2 D(f~p_2)a_1\texttt{;} \\ {\Delta}p_1 &=~~~ -\eta {{\hskip0.5mm\mathsf{d}}}_1 D(\lambda v.f~v~x) p_1\texttt{;} \\ \end{array}\)
This can be instantiated to the particular cost function \(C\) and activation function \(f\) discussed above, namely:
\(C~u~v = \frac{1}{2}(u - v)^2\)
\(f~w~a = \sigma(w a + b)\)
Thus from the calculation of derivatives above:
\(D(C~u) = D(\lambda v. \frac{1}{2}(u - v)^2) = \lambda v. v - u\)
\(D(f~w) = D(\lambda v. \sigma(w v + b)) = \lambda v. \sigma'(w v + b) w\)
\(D(\lambda v. f~v~a) = D(\lambda v. \sigma(v a + b)) = \lambda v. \sigma'(v a + b) a\)
The last equation above is derived from the preceding one as \(a\hskip0.25mm v = v\hskip0.25mm a\).
Taking the parameter \(p_i\) to be weight \(w_i\), the pseudocode thus instantiates to
Forward pass:
\(~~~\begin{array}{ll} a_1 &=~~~ \sigma(w_1 x + b)\texttt{;} \\ a_2 &=~~~ \sigma(w_2 a_1 + b)\texttt{;} \\ a_3 &=~~~ \sigma(w_3 a_2 + b)\texttt{;} \\ a_4 &=~~~ \sigma(w_4 a_b + b)\texttt{;} \\ \end{array}\)
Backward pass:
\(~~~\begin{array}{ll} {{\hskip0.5mm\mathsf{d}}}_4 &=~~~ a_4 - g~x\texttt{;} \\ {\Delta}w_4 &=~~~ -\eta {{\hskip0.5mm\mathsf{d}}}_4 \sigma'(w_4 a_3 + b) a_3 \texttt{;} \\ {{\hskip0.5mm\mathsf{d}}}_3 &=~~~ {{\hskip0.5mm\mathsf{d}}}_4 \sigma'(w_4 a_3 + b) w_4 \texttt{;} \\ {\Delta}w_3 &=~~~ -\eta {{\hskip0.5mm\mathsf{d}}}_3 \sigma'(w_3 a_2 + b) a_2 \texttt{;} \\ {{\hskip0.5mm\mathsf{d}}}_2 &=~~~ {{\hskip0.5mm\mathsf{d}}}_3 \sigma'(w_3 a_2 + b) w_3 \texttt{;} \\ {\Delta}w_2 &=~~~ -\eta {{\hskip0.5mm\mathsf{d}}}_2 \sigma'(w_2 a_1 + b) a_1 \texttt{;} \\ {{\hskip0.5mm\mathsf{d}}}_1 &=~~~ {{\hskip0.5mm\mathsf{d}}}_2 \sigma'(w_2 a_1 + b) w_2 \texttt{;} \\ {\Delta}w_1 &=~~~ -\eta {{\hskip0.5mm\mathsf{d}}}_1 \sigma'(w_1 x + b) x \texttt{;} \\ \end{array}\)
Unwinding the equations for the deltas:
\(~~~\begin{array}{l} {{\hskip0.5mm\mathsf{d}}}_4 = a_4 {-} g~x\texttt{;} \\ {\Delta}w_4 = {-}\eta (a_4 {-} g~x) \sigma'(w_4 a_3 {+} b) a_3 \texttt{;} \\ {{\hskip0.5mm\mathsf{d}}}_3 = (a_4 {-} g~x) \sigma'(w_4 a_3 {+} b) w_4 \texttt{;} \\ {\Delta}w_3 = {-}\eta (a_4 {-} g~x) \sigma'(w_4 a_3 {+} b) w_4 \sigma'(w_3 a_2 {+} b) a_2 \texttt{;} \\ {{\hskip0.5mm\mathsf{d}}}_2 = (a_4 {-} g~x) \sigma'(w_4 a_3 {+} b) w_4 \sigma'(w_3 a_2 {+} b) w_3 \texttt{;} \\ {\Delta}w_2 = {-}\eta (a_4 {-} g~x) \sigma'(w_4 a_3 {+} b) w_3 \sigma'(w_3 a_2 {+} b) w_2 \sigma'(w_2 a_1 {+} b) a_1 \texttt{;} \\ {{\hskip0.5mm\mathsf{d}}}_1 = (a_4 {-} g~x) \sigma'(w_4 a_3 {+} b) w_4 \sigma'(w_3 a_2 {+} b) w_3 \sigma'(w_2 a_1 {+} b) w_1 \texttt{;} \\ {\Delta}w_1 = {-}\eta (a_4 {-} g~x) \sigma'(w_4 a_3 {+} b) w_4 \sigma'(w_3 a_2 {+} b) w_3 \sigma'(w_2 a_1 {+} b) w_2 \sigma'(w_1 x {+} b) x \texttt{;} \\ \end{array}\)
So far each activation function has the same fixed bias. In practice each neuron has its own bias and these, like the weights, are learnt using gradient descent. This is shown in the diagram below where the weights are on the incoming arrows and the biases inside the nodes.
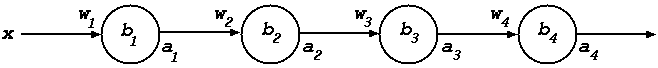
The calculation of the bias deltas is similar to – in fact simpler – than the calculation of the weight deltas.
Assume now that the activation function \(f\) has two real number parameters: a weight \(w\) and bias \(b\), so \(f : {\Bbb R}\rightarrow{\Bbb R}\rightarrow{\Bbb R}\).
To calculate the partial derivative of the activation function with respect to the bias let
\(a = f~w~b~v = \sigma(w v + b)\)
To compute \(\frac{\partial a}{\partial b}\) treat \(w\) and \(v\) as constants, then use the chain rule to derive \(\frac{da}{db}\).
\(\begin{array}{l|l|l} \hline {\textbf{Equation}} & {\textbf{Derivative}} & {\textbf{Differentiation rules used}} \\ \hline a = \sigma({z}) & \frac{da}{d{z}} = \sigma({z})(1{-}\sigma({z})) & \textrm{see discussion above} \\ \hline {z} = w v + b & \frac{d{z}}{db} = 1 & \textrm{addition, constant}, \frac{db}{db}{=}1\\ \hline \end{array}\)
Hence \(\frac{da}{db} = \frac{da}{d{z}} \times \frac{d{z}}{db} = \sigma({z})(1{-}\sigma({z})) \times 1 = \sigma(w v + b)(1{-}\sigma(w v + b)) \times 1\).
Thus \(\frac{\partial a}{\partial b} = \sigma'(w v + b)\). By an earlier calculation \(\frac{\partial a}{\partial v} = \sigma'(w v + b)w\).
The activations are:
\(~~~\begin{array}{ll} a_1 &=~~~ \sigma(w_1 x + b_1) \\ a_2 &=~~~ \sigma(w_2 a_1 + b_2) \\ a_3 &=~~~ \sigma(w_3 a_2 + b_3) \\ a_4 &=~~~ \sigma(w_4 a_3 + b_4) \\ \end{array}\)
The cost function equation is:
\(c = C(g~x)(f~w_4~b_4(f~w_3~b_3(f~w_2~b_2(f~w_1~b_1~x)))) = C (g~x) a_4\)
The bias deltas are \({\Delta}b_i = -\eta\hskip1mm \frac{\partial c}{\partial b_i}\). The argument for this is similar to the argument for the weight deltas.
By the chain rule:
\(\frac{\partial c}{\partial b_1} = \frac{\partial c}{\partial a_4} \frac{\partial a_4}{\partial a_3} \frac{\partial a_3}{\partial a_2} \frac{\partial a_2}{\partial a_1} \frac{\partial a_1}{\partial b_1}\)
\(\frac{\partial c}{\partial b_2} = \frac{\partial c}{\partial a_4} \frac{\partial a_4}{\partial a_3} \frac{\partial a_3}{\partial a_2} \frac{\partial a_2}{\partial b_2}\)
\(\frac{\partial c}{\partial b_3} = \frac{\partial c}{\partial a_4} \frac{\partial a_4}{\partial a_3} \frac{\partial a_3}{\partial b_3}\)
\(\frac{\partial c}{\partial b_4} = \frac{\partial c}{\partial a_4} \frac{\partial a_4}{\partial b_4}\)
Calculate as follows:
\({{\hskip0.5mm\mathsf{d}}}_4 = \frac{\partial c}{\partial a_4} = a_4 - g~x\)
\({\Delta}b_4 = - \eta\hskip1mm \frac{\partial c}{\partial a_4} \frac{\partial a_4}{\partial b_4}\\ \phantom{{\Delta}b_4} = - \eta\hskip1mm {{\hskip0.5mm\mathsf{d}}}_4 \sigma'(w_4 a_3 + b_4)\)
\({{\hskip0.5mm\mathsf{d}}}_3 = \frac{\partial c}{\partial a_4} \frac{\partial a_4}{\partial a_3} = {{\hskip0.5mm\mathsf{d}}}_4 \sigma'(w_4 a_3 + b_4)w_4\)
\({\Delta}b_3 = - \eta\hskip1mm \frac{\partial c}{\partial a_4} \frac{\partial a_4}{\partial a_3} \frac{\partial a_3}{\partial b_3}\\ \phantom{{\Delta}b_3} = - \eta\hskip1mm {{\hskip0.5mm\mathsf{d}}}_3 \sigma'(w_3 a_2 + b_3)\)
\({{\hskip0.5mm\mathsf{d}}}_2 = \frac{\partial c}{\partial a_4} \frac{\partial a_4}{\partial a_3} \frac{\partial a_3}{\partial a_2} = {{\hskip0.5mm\mathsf{d}}}_3 \sigma'(w_3 a_2 + b_3) w_3\)
\({\Delta}b_2 = - \eta\hskip1mm \frac{\partial c}{\partial a_4} \frac{\partial a_4}{\partial a_3} \frac{\partial a_3}{\partial a_2} \frac{\partial a_2}{\partial b_2}\\ \phantom{{\Delta}b_2} = - \eta\hskip1mm {{\hskip0.5mm\mathsf{d}}}_2 \sigma'(w_2 a_1 + b_2)\)
\({{\hskip0.5mm\mathsf{d}}}_1 = \frac{\partial c}{\partial a_4} \frac{\partial a_4}{\partial a_3} \frac{\partial a_3}{\partial a_2} \frac{\partial a_2}{\partial a_1} = {{\hskip0.5mm\mathsf{d}}}_2 \sigma'(w_2 a_1 + b_2) w_2\)
\({\Delta}b_1 = - \eta\hskip1mm \frac{\partial c}{\partial a_4} \frac{\partial a_4}{\partial a_3} \frac{\partial a_3}{\partial a_2} \frac{\partial a_2}{\partial a_1} \frac{\partial a_1}{\partial b_1}\\ \phantom{{\Delta}b_1} = - \eta\hskip1mm {{\hskip0.5mm\mathsf{d}}}_1 \sigma'(w_1 x + b_1)\)
Summarising the bias deltas calculaton:
\(\begin{array}{ll} {{\hskip0.5mm\mathsf{d}}}_4 = a_4 - g~x \qquad\qquad & {\Delta}b_4 = - \eta\hskip1mm {{\hskip0.5mm\mathsf{d}}}_4 \sigma'(w_4 a_3 + b_4)\\ {{\hskip0.5mm\mathsf{d}}}_3 = {{\hskip0.5mm\mathsf{d}}}_4 \sigma'(w_4 a_3 + b_4)w_4& {\Delta}b_3 = - \eta\hskip1mm {{\hskip0.5mm\mathsf{d}}}_3 \sigma'(w_3 a_2 + b_3)\\ {{\hskip0.5mm\mathsf{d}}}_2 = {{\hskip0.5mm\mathsf{d}}}_3 \sigma'(w_3 a_2 + b_3) w_3 & {\Delta}b_2 = - \eta\hskip1mm {{\hskip0.5mm\mathsf{d}}}_2 \sigma'(w_2 a_1 + b_2)\\ {{\hskip0.5mm\mathsf{d}}}_1 = {{\hskip0.5mm\mathsf{d}}}_2 \sigma'(w_2 a_1 + b_2) w_2 & {\Delta}b_1 = - \eta\hskip1mm {{\hskip0.5mm\mathsf{d}}}_1 \sigma'(w_1 x + b_1)\\ \end{array}\)
The weight delta calculations with separate biases for each neuron are produced by tweaking the calculation in the previous section.
\(\begin{array}{ll} {{\hskip0.5mm\mathsf{d}}}_4 = a_4 - g~x \qquad\qquad & {\Delta}w_4 = - \eta\hskip1mm {{\hskip0.5mm\mathsf{d}}}_4 \sigma'(w_4 a_3 + b_4) a_3 \\ {{\hskip0.5mm\mathsf{d}}}_3 = {{\hskip0.5mm\mathsf{d}}}_4 \sigma'(w_4 a_3 + b_4)w_4& {\Delta}w_3 = - \eta\hskip1mm {{\hskip0.5mm\mathsf{d}}}_3 \sigma'(w_3 a_2 + b_3) a_2 \\ {{\hskip0.5mm\mathsf{d}}}_2 = {{\hskip0.5mm\mathsf{d}}}_3 \sigma'(w_3 a_2 + b_3) w_3 & {\Delta}w_2 = - \eta\hskip1mm {{\hskip0.5mm\mathsf{d}}}_2 \sigma'(w_2 a_1 + b_2) a_1 \\ {{\hskip0.5mm\mathsf{d}}}_1 = {{\hskip0.5mm\mathsf{d}}}_2 \sigma'(w_2 a_1 + b_2) w_2 & {\Delta}w_1 = - \eta\hskip1mm {{\hskip0.5mm\mathsf{d}}}_1 \sigma'(w_1 x + b_1) x \\ \end{array}\)
If \(\delta_i = {{\hskip0.5mm\mathsf{d}}}_i \sigma'(w_i a_{i{-}1} + b_i)\) then one can summarise everything by:
\(\begin{array}{lll} \delta_4 = (a_4 - g~x)\sigma'(w_4 a_3 + b_4) & {\Delta}w_4 = - \eta\hskip1mm \delta_4 a_3 & {\Delta}b_4 = - \eta\hskip1mm \delta_4 \\ \delta_3 = \delta_4 \sigma'(w_3 a_2 + b_3) w_4 & {\Delta}w_3 = - \eta\hskip1mm \delta_3 a_2 & {\Delta}b_3 = - \eta\hskip1mm \delta_3 \\ \delta_2 = \delta_3 \sigma'(w_2 a_1 + b_2) w_3 & {\Delta}w_2 = - \eta\hskip1mm \delta_2 a_1 & {\Delta}b_2 = - \eta\hskip1mm \delta_2 \\ \delta_1 = \delta_2 \sigma'(w_1 x + b_1) w_2 & {\Delta}w_1 = - \eta\hskip1mm \delta_1 x & {\Delta}b_1 = - \eta\hskip1mm \delta_1 \\ \end{array}\)
Consider now a network that uses the logistic \(\sigma\) function and has four layers and two neurons per layer.
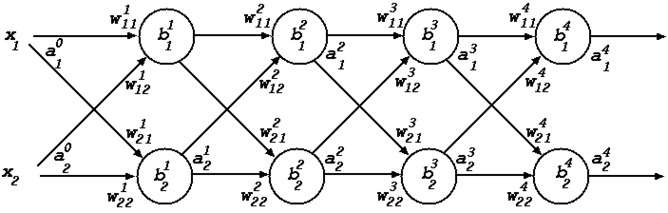
The notation from Chapter 2 of Nielsen’s book is used here. Note that superscripts are used to specify the layer that a variable corresponds to. Up until now – e.g. in the one-neuron-per-layer example above – subscripts have been used for this. The notation below uses subscripts to specify which neuron in a layer a variable is associated with.
\(L\) is the number of layers; \(L=4\) in the example.
\(y_j\) is the desired output from the \(j^{\textrm{th}}\) neuron; if the network has \(r\) inputs and \(s\) ouputs and \(g : {\Bbb R}^r\rightarrow{\Bbb R}^s\) is the function the network is intended to approximate, then \((y_1,\ldots,y_s) = g(x_1,\ldots,x_r)\) (\(r=s=2\) in the example here).
\(a^0_j\) is the \(j^{\textrm{th}}\) input, also called \(x_j\); if \(l>0\) then \(a^l_j\) is the output, i.e. the activation, from the \(j^{\textrm{th}}\) neuron in the \(l^{\textrm{th}}\) layer.
\(w^l_{jk}\) is the weight from the \(k^{\textrm{th}}\) neuron in the \((l{-}1)^{\textrm{th}}\) layer to the \(j^{\textrm{th}}\) neuron in the \(l^{\textrm{th}}\) layer (the reason for this ordering of subscripts – “\(jk\)” rather than the apparently more natural “\(kj\)” – is explained in Chapter 2 of Nielsen’s book: the chosen order is to avoid a matrix transpose later on; for more details search the web page for “quirk”).
\(b^l_j\) is the bias for the \(j^{\textrm{th}}\) neuron in the \(l^{\textrm{th}}\) layer.
\(z^l_j\) is the weighted input to the sigmoid function for the \(j^{\textrm{th}}\) neuron in the \(l^{\textrm{th}}\) layer: \(z^l_j = (\sum\limits_k w^l_{jk} a^{l{-}1}_k) + b^l_j\) – thus \(a^l_j = \sigma(z^l_j) = \sigma(\sum\limits_k w^l_{jk} a^{l{-}1}_k + b^l_j)\).
Note that \(z^L_j\) are not the outputs of the network. I mention this to avoid confusion, as \(z\) was used for the output back in the section entitled Gradient descent. With the variable name conventions here, the outputs are \(a^L_j\).
For the network in the diagram \(k\in\{1,2\}\), so \(a^l_j = \sigma(z^l_j) = \sigma(w^l_{j1} a^{l{-}1}_1 + w^l_{j2} a^{l{-}1}_2 + b^l_j)\).
For more than one output, the mean square error cost function \(C\) is
\(C (u_1,\ldots,u_s) (v_1,\ldots,v_s) = \frac{1}{2} \left\| (u_1,\ldots,u_s) - (v_1,\ldots,v_s)\right\|^{2}\)
where \(\left\|(u_1,\ldots,u_s)\right\| = \sqrt{u_1^2 + \cdots + u_s^2}\) is the norm of the vector \((u_1,\ldots,u_s)\). The norm is also sometimes called the length, but there is potential confusion with “length” also meaning the number of components or dimension.
Thus
\(\begin{array}{l} C (u_1,\ldots,u_s) (v_1,\ldots,v_s) \\ ~ = \frac{1}{2} \left\| (u_1,\ldots,u_s) - (v_1,\ldots,v_s)\right\|^{2}\\ ~ = \frac{1}{2} \left\| ((u_1-v_1),\ldots,(u_s-v_s))\right\|^{2}\\ ~ = \frac{1}{2} \left(\hskip1mm\sqrt{(u_1-v_1)^2+\cdots+(u_s-v_s)^2}\hskip1mm \right)^2\\ ~ = \frac{1}{2} ((v_1 - u_1)^2 + \cdots + (v_s - u_s)^2)\\ \end{array}\)
If the network computes function \(f_{\scriptsize\textsf{N}}\), which will depend on the values of all the weights \(w^l_{jk}\) and biases \(b^l_j\), then for a given input vector \((x_1,\ldots,x_r) = (a^0_1,\ldots,a^0_r)\), the cost to be minimised is
\(\begin{array}{l}C (g(x_1,\ldots,x_r))(f_{\scriptsize\textsf{N}}(x_1,\ldots,x_r))\\ ~ = \frac{1}{2} \left\|g(x_1,\ldots,x_r) - f_{\scriptsize\textsf{N}}(x_1,\ldots,x_r) \right\|^{2}\\ ~ = \frac{1}{2} \left\|(y_1,\ldots,y_s) - (a^L_1,\ldots,a^L_s) \right\|^{2}\\ ~ = \frac{1}{2} ((y_1 - a^L_1)^2 + \cdots + (y_s - a^L_s)^2)\\ \end{array}\)
For the two neurons per layer example this becomes: \(C(g(x_1,x_2))(f_{\scriptsize\textsf{N}}(x_1,x_2)) = \frac{1}{2} \left\|g(x_1,x_2) - f_{\scriptsize\textsf{N}}(x_1,x_2)\right\|^{2} = \frac{1}{2} ((y_1{-}a^4_1)^2 + (y_2{-}a^4_2)^2)\).
The activations of the first layer are shown below both as a pair or equations and as a single equation using vector and matrix addition and multiplication. The function \(\sigma\) is interpreted as acting elementwise on vectors – i.e. the application of \(\sigma\) to a vector is interpreted as the vector whose components are the result of applying \(\sigma\) to each of them.
\(\begin{array}{lr} \begin{array}{l} a^1_1 = \sigma(w^1_{11} a^0_1 + w^1_{12} a^0_2 + b^1_1)\\ a^1_2 = \sigma(w^1_{21} a^0_1 + w^1_{22} a^0_2 + b^1_2)\\ \end{array} & ~~~~ \left\lbrack \begin{matrix} a^1_1 \cr a^1_2 \cr \end{matrix}\right\rbrack = \sigma\left( \left\lbrack \begin{matrix} w^1_{11} & w^1_{12} \cr w^1_{21} & w^1_{22} \cr \end{matrix}\right\rbrack {\times} \left\lbrack \begin{matrix} a^0_1 \cr a^0_2 \cr \end{matrix}\right\rbrack {+} \left\lbrack \begin{matrix} b^1_1 \cr b^1_2 \cr \end{matrix}\right\rbrack\right) \end{array}\)
Similarly for the layer 2 activations.
\(\begin{array}{lr} \begin{array}{l} a^2_1 = \sigma(w^2_{11} a^1_1 + w^2_{12} a^1_2 + b^2_1)\\ a^2_2 = \sigma(w^2_{21} a^1_1 + w^2_{22} a^1_2 + b^2_2)\\ \end{array} & ~~~~ \left\lbrack \begin{matrix} a^2_1 \cr a^2_2 \cr \end{matrix}\right\rbrack = \sigma\left( \left\lbrack \begin{matrix} w^2_{11} & w^2_{12} \cr w^2_{21} & w^2_{22} \cr \end{matrix}\right\rbrack {\times} \left\lbrack \begin{matrix} a^1_1 \cr a^1_2 \cr \end{matrix}\right\rbrack {+} \left\lbrack \begin{matrix} b^2_1 \cr b^2_2 \cr \end{matrix}\right\rbrack\right) \end{array}\)
These show the general pattern.
\(\begin{array}{lr} \begin{array}{l} a^l_1 = w^l_{11} a^{(l-1)}_1 + w^l_{12} a^{(l-1)}_2 + b^l_1\\ a^l_2 = w^l_{21} a^{(l-1)}_1 + w^l_{22} a^{(l-1)}_2 + b^l_2\\ \end{array} & \left\lbrack \begin{matrix} a^l_1 \cr a^l_2 \cr \end{matrix}\right\rbrack = \sigma\left( \left\lbrack \begin{matrix} w^l_{11} & w^l_{12} \cr w^l_{21} & w^l_{22} \cr \end{matrix}\right\rbrack {\times} \left\lbrack \begin{matrix} a^{(l-1)}_1 \cr a^{(l-1)}_2 \cr \end{matrix}\right\rbrack {+} \left\lbrack \begin{matrix} b^l_1 \cr b^l_2 \cr \end{matrix}\right\rbrack\right) \end{array}\)
This can be written as a vector equation \(a^l = \sigma(w^l \times a^{(l-1)} + b^l)\) where:
\(a^l = \left\lbrack \begin{matrix} a^l_1 \cr a^l_2 \cr \end{matrix}\right\rbrack\) \(\textrm{and}~\) \(w^l = \left\lbrack \begin{matrix} w^l_{11} & w^l_{12} \cr w^l_{21} & w^l_{22} \cr \end{matrix}\right\rbrack\) \(\textrm{and}~\) \(a^{(l-1)} = \left\lbrack \begin{matrix} a^{(l-1)}_1 \cr a^{(l-1)}_2 \cr \end{matrix}\right\rbrack\) \(\textrm{and}~\) \(b^l = \left\lbrack \begin{matrix} b^l_1 \cr b^l_2 \cr \end{matrix}\right\rbrack\).
In the one-neuron-per-layer example, each activation is a single real number and is entirely determined by the activation of the preceding layer. The actual output of the network is \(a_4\) and the desired output, \(y\) say, is \(g~x\). The error \(c\) that it is hoped to reduce by learning values of the weights and biases is:
\(c = C (g~x)(a_4) = \frac{1}{2}(y - a_4)^2\)
Thus \(c\) depends on just one output, \(a_4\) and \(\frac{\partial c}{\partial w_i} = \frac{\partial c}{\partial a_4} \frac{\partial a_4}{\partial w_i}\).
To avoid confusion here, remember that in the one-neuron-per-layer net example the layer of weights and biases is specified with a subscript, e.g. \(a_4\) is the activation from layer 4, but in the two-neuron-per-layer example these are specified by a superscript, e.g. \(a^4_1\) is the activation from neuron 1 in layer 4.
In the two-neuron-per-layer example: \(c = \frac{1}{2} ((y_1 - a^4_1)^2 + (y_2 - a^4_2)^2)\) and \(c\) depends on two variables: \(a^4_1\) and \(a^4_2\). The contribution of changes in a weight \(w^l_{jk}\) to changes in \(c\) is the combination of the changes it makes to the two outputs \(a^4_1\) and \(a^4_2\). How these contributions combine is specified by the chain rule, which states that if a variable \(u\) depends on variables \(v_1\),\(\ldots\), \(v_n\) and if each \(v_i\) depends on \(t\), then
\(\frac{du}{dt} = \frac{du}{dv_1}\frac{dv_1}{dt} + \cdots + \frac{du}{dv_n}\frac{dv_n}{dt}\)
The “\(\scriptsize{d}\)” becomes “\(\scriptsize{\partial}\)” if there are other variables that are being treated as constants.
The error \(c\) depends on both \(a^4_1\) and \(a^4_2\), which in turn depend on the weights \(w^l_{jk}\) and biases \(b^l_j\), hence \(\frac{\partial c}{\partial w^l_{jk}}\) and \(\frac{\partial c}{\partial b^l_j}\). Applying the chain rule to the example gives:
\(\begin{array}{ll} \frac{\partial c}{\partial w^l_{jk}} = \frac{\partial c}{\partial a^4_1} \frac{\partial a^4_1}{\partial w^l_{jk}} + \frac{\partial c}{\partial a^4_2} \frac{\partial a^4_2}{\partial w^l_{jk}} &\qquad \frac{\partial c}{\partial b^l_j} = \frac{\partial c}{\partial a^4_1} \frac{\partial a^4_1}{\partial b^l_j} + \frac{\partial c}{\partial a^4_2} \frac{\partial a^4_2}{\partial b^l_j}\\ \frac{\partial c}{\partial w^l_{jk}} = \frac{\partial c}{\partial a^4_1} \frac{\partial a^4_1}{\partial z^4_1} \frac{\partial z^4_1}{\partial w^l_{jk}} + \frac{\partial c}{\partial a^4_2} \frac{\partial a^4_2}{\partial z^4_2} \frac{\partial z^4_2}{\partial w^l_{jk}} &\qquad \frac{\partial c}{\partial b^l_j} = \frac{\partial c}{\partial a^4_1} \frac{\partial a^4_1}{\partial z^4_1} \frac{\partial z^4_1}{\partial b^l_j} + \frac{\partial c}{\partial a^4_2} \frac{\partial a^4_2}{\partial z^4_1} \frac{\partial z^4_2}{\partial b^l_j} \end{array}\)
More generally, if there are many neurons per layer then the derivatives from each layer are summed:
\(\frac{\partial c}{\partial w^l_{jk}} = \sum\limits_i \frac{\partial c}{\partial a^L_i} \frac{\partial a^L_i}{\partial w^l_{jk}}\) \(\qquad\) \(\frac{\partial c}{\partial b^l_j} = \sum\limits_i \frac{\partial c}{\partial a^L_i} \frac{\partial a^L_i}{\partial b^l_i}\)
Return now to the two-neuron-per-layer example, where \(i=1\) or \(i=2\), \(L=4\) and the cost is \(c = \frac{1}{2} ((y_1 - a^4_1)^2 + (y_2 - a^4_2)^2)\) which can be split into \(c~=~\frac{1}{2}(u{+}(y_2{-}a^4_2)^2)\) \(~\textrm{and}~\) \(u~=~v^2\) \(~\textrm{and}~\) \(v~=~(y_1{-}a^4_1)\). By the chain rule
\(\frac{\partial c}{\partial a^4_1} = \frac{\partial c}{\partial u} {\times} \frac{\partial u}{\partial v} {\times} \frac{\partial v}{\partial a^4_1} = \frac{1}{2} {\times} 2v {\times} (0 - 1) = \frac{1}{2} {\times} 2v {\times} {-}1 = {-}v = {-}(y_1 - a^4_1)\)
and similarly \(\frac{\partial c}{\partial a^4_2} = {-}(y_2 - a^4_2)\). To summarise (and also simplifying):
\(\frac{\partial c}{\partial a^4_1} = (a^4_1 - y_1)\) \(\qquad\) \(\frac{\partial c}{\partial a^4_2} = (a^4_2 - y_2)\)
The derivatives of \(c\) with respect to the last layer weights and biases are:
\(\frac{\partial c}{\partial w^4_{11}} = \frac{\partial c}{\partial a^4_1} \frac{\partial a^4_1}{\partial w^4_{11}} + \frac{\partial c}{\partial a^4_2} \frac{\partial a^4_2}{\partial w^4_{11}}\) \(\qquad\) \(\frac{\partial c}{\partial b^4_1} = \frac{\partial c}{\partial a^4_1} \frac{\partial a^4_1}{\partial b^4_1} + \frac{\partial c}{\partial a^4_2} \frac{\partial a^4_2}{\partial b^4_1}\)
\(\frac{\partial c}{\partial w^4_{12}} = \frac{\partial c}{\partial a^4_1} \frac{\partial a^4_1}{\partial w^4_{12}} + \frac{\partial c}{\partial a^4_2} \frac{\partial a^4_2}{\partial w^4_{12}}\) \(\qquad\) \(\frac{\partial c}{\partial b^4_2} = \frac{\partial c}{\partial a^4_1} \frac{\partial a^4_1}{\partial b^4_2} + \frac{\partial c}{\partial a^4_2} \frac{\partial a^4_2}{\partial b^4_2}\)
In order to use the chain rule, it’s convenient to express the equations for the activations \(a^l_j\) using the weighted inputs \(z^l_j\).
\(\begin{array}{ll} a^1_1 = \sigma(z^1_1) & \qquad z^1_1 = w^1_{11} a^0_1 + w^1_{12} a^0_2 + b^1_1\\ a^1_2 = \sigma(z^1_2) & \qquad z^1_2 = w^1_{21} a^0_1 + w^1_{22} a^0_2 + b^1_2\\ \end{array}\)
\(\begin{array}{ll} a^2_1 = \sigma(z^2_1) & \qquad z^2_1 = w^2_{11} a^1_1 + w^2_{12} a^1_2 + b^2_1\\ a^2_2 = \sigma(z^2_2) & \qquad z^2_2 = w^2_{21} a^1_1 + w^2_{22} a^1_2 + b^2_2\\ \end{array}\)
\(\begin{array}{ll} a^3_1 = \sigma(z^3_1) & \qquad z^3_1 = w^3_{11} a^2_1 + w^3_{12} a^2_2 + b^3_1\\ a^3_2 = \sigma(z^3_2) & \qquad z^3_2 = w^3_{21} a^2_1 + w^3_{22} a^2_2 + b^3_2\\ \end{array}\)
\(\begin{array}{ll} a^4_1 = \sigma(z^4_1) & \qquad z^4_1 = w^4_{11} a^3_1 + w^4_{12} a^3_2 + b^4_1\\ a^4_2 = \sigma(z^4_2) & \qquad z^4_2 = w^4_{21} a^3_1 + w^4_{22} a^3_2 + b^4_2\\ \end{array}\)
It turns out, as nicely explained in Chapter 2 of Nielsen’s book, that things work out especially neatly if one formulates the calculations of the weight and bias deltas in terms of \(\delta^l_j = \frac{\partial c}{\partial z^l_j}\). The calculations proceed backwards.
For the last layer of the example:
\(\begin{array}{l} \delta^4_1 = \frac{\partial c}{\partial z^4_1} = \frac{\partial c}{\partial a^4_1} \frac{\partial a^4_1}{\partial z^4_1} = (a^4_1 - y_1) \sigma'(z^4_1)\\ \end{array}\)
\(\begin{array}{l} \delta^4_2 = \frac{\partial c}{\partial z^4_2} = \frac{\partial c}{\partial a^4_2} \frac{\partial a^4_2}{\partial z^4_2} = (a^4_2 - y_2) \sigma'(z^4_2)\\ \end{array}\)
The derivatives of \(c\) with respect to the weights in the last layer are expressed in terms of \(\delta^4_1\) and \(\delta^4_2\) below. First note that:
\(\begin{array}{l} c = \frac{1}{2} ((y_1 - \sigma(z^4_1))^2 + (y_2 - \sigma(z^4_2))^2)\\ \phantom{c} = \frac{1}{2} ((y_1 - \sigma(z^4_1))^2 + (y_2 - \sigma(z^4_2))^2) \end{array}\)
By the chain rule
\(\begin{array}{l} \frac{\partial c}{\partial w^4_{11}} = \frac{\partial c}{\partial z^4_1} \frac{\partial z^4_1}{\partial w^4_{11}} + \frac{\partial c}{\partial z^4_2} \frac{\partial z^4_2}{\partial w^4_{11}} \\ \phantom{\frac{\partial c}{\partial w^4_{11}}} = \delta^4_1 \frac{\partial z^4_1}{\partial w^4_{11}} + \delta^4_2 \frac{\partial z^4_2}{\partial w^4_{11}} \\ \end{array}\)
\(\begin{array}{l} \frac{\partial c}{\partial w^4_{12}} = \frac{\partial c}{\partial z^4_1} \frac{\partial z^4_1}{\partial w^4_{12}} + \frac{\partial c}{\partial z^4_2} \frac{\partial z^4_2}{\partial w^4_{12}} \\ \phantom{\frac{\partial c}{\partial w^4_{12}}} = \delta^4_1 \frac{\partial z^4_1}{\partial w^4_{12}} + \delta^4_2 \frac{\partial z^4_2}{\partial w^4_{12}} \\ \end{array}\)
\(\begin{array}{l} \frac{\partial c}{\partial w^4_{21}} = \frac{\partial c}{\partial z^4_1} \frac{\partial z^4_1}{\partial w^4_{21}} + \frac{\partial c}{\partial z^4_2} \frac{\partial z^4_2}{\partial w^4_{21}} \\ \phantom{\frac{\partial c}{\partial w^4_{21}}} = \delta^4_1 \frac{\partial z^4_1}{\partial w^4_{21}} + \delta^4_2 \frac{\partial z^4_2}{\partial w^4_{21}} \\ \end{array}\)
\(\begin{array}{l} \frac{\partial c}{\partial w^4_{22}} = \frac{\partial c}{\partial z^4_1} \frac{\partial z^4_1}{\partial w^4_{22}} + \frac{\partial c}{\partial z^4_2} \frac{\partial z^4_2}{\partial w^4_{22}} \\ \phantom{\frac{\partial c}{\partial w^4_{22}}} = \delta^4_1 \frac{\partial z^4_1}{\partial w^4_{22}} + \delta^4_2 \frac{\partial z^4_2}{\partial w^4_{22}} \\ \end{array}\)
and the derivatives of the biases are:
\(\begin{array}{l} \frac{\partial c}{\partial b^4_1} = \frac{\partial c}{\partial z^4_1} \frac{\partial z^4_1}{\partial b^4_1} + \frac{\partial c}{\partial z^4_2} \frac{\partial z^4_2}{\partial b^4_1}\\ \phantom{\frac{\partial c}{\partial b^4_1}} = \delta^4_1 \frac{\partial z^4_1}{\partial b^4_1} + \delta^4_2 \frac{\partial z^4_2}{\partial b^4_1}\\ \end{array}\)
\(\begin{array}{l} \frac{\partial c}{\partial b^4_2} = \frac{\partial c}{\partial z^4_1} \frac{\partial z^4_1}{\partial b^4_2} + \frac{\partial c}{\partial z^4_2} \frac{\partial z^4_2}{\partial b^4_2}\\ \phantom{\frac{\partial c}{\partial b^4_2}} = \delta^4_1 \frac{\partial z^4_1}{\partial b^4_2} + \delta^4_2 \frac{\partial z^4_2}{\partial b^4_2}\\ \end{array}\)
By the addition and multiplication-by-a-constant rules for differentiation
\(\frac{\partial z^4_j}{\partial w^4_{jk}} = a^3_k\) \(\qquad\) \(\frac{\partial z^4_j}{\partial b^4_j} = 1\)
and if \(i\neq j\) then:
\(\frac{\partial z^4_i}{\partial w^4_jk} = 0\) \(\qquad\) \(\frac{\partial z^4_i}{\partial b^4_j} = 0\)
Using these and simple arithmetic yields the following summary of the derivatives of \(c\) with respect to the weights and biases of the last layer.
\(\begin{array}{l} \frac{\partial c}{\partial w^4_{11}} = \delta^4_1 a^3_1 \end{array}\) \(\qquad\) \(\begin{array}{l} \frac{\partial c}{\partial w^4_{21}} = \delta^4_2 a^3_1 \\ \end{array}\) \(\qquad\) \(\begin{array}{l} \frac{\partial c}{\partial b^4_1} = \delta^4_1 \end{array}\)
\(\begin{array}{l} \frac{\partial c}{\partial w^4_{12}} = \delta^4_1 a^3_2 \end{array}\) \(\qquad\) \(\begin{array}{l} \frac{\partial c}{\partial w^4_{22}} = \delta^4_2 a^3_2 \\ \end{array}\) \(\qquad\) \(\begin{array}{l} \frac{\partial c}{\partial b^4_2} = \delta^4_2\\ \end{array}\)
expanding \(\delta^4_j\) gives:
\(\begin{array}{lll} \begin{array}{l} \frac{\partial c}{\partial w^4_{11}} = (a^4_1-y_1) \sigma'(z^4_1) a^3_1 \end{array} & \begin{array}{l} \frac{\partial c}{\partial w^4_{21}} = (a^4_2-y_2) \sigma'(z^4_2) a^3_1 \\ \end{array} & \begin{array}{l} \frac{\partial c}{\partial b^4_1} = (a^4_1-y_1) \sigma'(z^4_1) \end{array} \end{array}\)
\(\begin{array}{lll}\begin{array}{l} \frac{\partial c}{\partial w^4_{12}} = (a^4_1-y_1) \sigma'(z^4_1) a^3_2 \end{array} & \begin{array}{l} \frac{\partial c}{\partial w^4_{22}} = (a^4_2-y_2) \sigma'(z^4_2) a^3_2 \\ \end{array} & \begin{array}{l} \frac{\partial c}{\partial b^4_2} = (a^4_2-y_2) \sigma'(z^4_2)\\ \end{array} \end{array}\)
Now consider the derivatives with respect to the weights of the top neurons in layer 3 of the example: \(\frac{\partial z^3_j}{\partial w^3_{jk}} = a^2_k\) and \(\frac{\partial z^3_j}{\partial b^3_j} = 1\), and if \(i\neq j\) then \(\frac{\partial z^3_i}{\partial w^3_jk} = 0\) and \(\frac{\partial z^3_i}{\partial b^3_j} = 0\).
Also:
\(\begin{array}{l} \delta^3_1 = \frac{\partial c}{\partial z^3_1}\\ \phantom{\delta^3_1} = \frac{\partial c}{\partial z^4_1} \frac{\partial z^4_1}{\partial a^3_1} \frac{\partial a^3_1}{\partial z^3_1} + \frac{\partial c}{\partial z^4_2} \frac{\partial z^4_2}{\partial a^3_1} \frac{\partial a^3_1}{\partial z^3_1}\\ \phantom{\delta^3_1} = \delta^4_1 w^4_{11} \sigma'(z^3_1) + \delta^4_2 w^4_{21} \sigma'(z^3_1)\\ \phantom{\delta^3_1} = (\delta^4_1 w^4_{11} + \delta^4_2 w^4_{21}) \sigma'(z^3_1)\\ \end{array}\) \(\qquad\) \(\begin{array}{l} \delta^3_2 = \frac{\partial c}{\partial z^3_2}\\ \phantom{\delta^3_2} = \frac{\partial c}{\partial z^4_1} \frac{\partial z^4_1}{\partial a^3_2} \frac{\partial a^3_2}{\partial z^3_2} + \frac{\partial c}{\partial z^4_2} \frac{\partial z^4_2}{\partial a^3_2} \frac{\partial a^3_2}{\partial z^3_2}\\ \phantom{\delta^3_2} = \delta^4_1 w^4_{12} \sigma(z^3_2) + \delta^4_2 w^4_{22} \sigma'(z^3_2)\\ \phantom{\delta^3_2} = (\delta^4_1 w^4_{12} + \delta^4_2 w^4_{22}) \sigma'(z^3_2)\\ \end{array}\)
Now
\(\begin{array}{l} \frac{\partial c}{\partial w^3_{11}} = \frac{\partial c}{\partial a^4_1} \frac{\partial a^4_1}{\partial w^3_{11}} + \frac{\partial c}{\partial a^4_2} \frac{\partial a^4_2}{\partial w^3_{11}} \\ \phantom{\frac{\partial c}{\partial w^4_{11}}} = \frac{\partial c}{\partial a^4_1} \frac{\partial a^4_1}{\partial z^4_1} \frac{\partial z^4_1}{\partial w^3_{11}} + \frac{\partial c}{\partial a^4_2} \frac{\partial a^4_2}{\partial z^4_2} \frac{\partial z^4_2}{\partial w^3_{11}} \\ \phantom{\frac{\partial c}{\partial w^4_{11}}} = \delta^4_1 \frac{\partial z^4_1}{\partial w^3_{11}} + \delta^4_2 \frac{\partial z^4_2}{\partial w^3_{11}} \\ \end{array}\)
and
\(\begin{array}{l} \frac{\partial z^4_1}{\partial w^3_{11}} = \frac{\partial z^4_1}{\partial a^3_1} \frac{\partial a^3_1}{\partial z^3_1} \frac{\partial z^3_1}{\partial w^3_{11}} + \frac{\partial z^4_1}{\partial a^3_2} \frac{\partial a^3_2}{\partial z^3_2} \frac{\partial z^3_2}{\partial w^3_{11}}\\ \phantom{\frac{\partial z^4_1}{\partial w^3_{11}}} = w^4_{11}{\times} \sigma'(z^3_1){\times} a^2_1 + w^4_{12}{\times} \sigma'(z^3_2){\times} 0\\ \phantom{\frac{\partial z^4_1}{\partial w^3_{11}}} = w^4_{11}{\times} \sigma'(z^3_1){\times} a^2_1\\ \end{array}\)
and
\(\begin{array}{l} \frac{\partial z^4_2}{\partial w^3_{11}} = \frac{\partial z^4_2}{\partial a^3_1} \frac{\partial a^3_1}{\partial z^3_1} \frac{\partial z^3_1}{\partial w^3_{11}} + \frac{\partial z^4_2}{\partial a^3_2} \frac{\partial a^3_2}{\partial z^3_2} \frac{\partial z^3_2}{\partial w^3_{11}}\\ \phantom{\frac{\partial z^4_1}{\partial w^3_{11}}} = w^4_{21}{\times} \sigma'(z^3_1){\times} a^2_1 + w^4_{22}{\times} \sigma'(z^3_2){\times} 0\\ \phantom{\frac{\partial z^4_1}{\partial w^3_{11}}} = w^4_{21}{\times} \sigma'(z^3_1){\times} a^2_1\\ \end{array}\)
Hence:
\(\begin{array}{l} \frac{\partial c}{\partial w^3_{11}} = \delta^4_1 \frac{\partial z^4_1}{\partial w^3_{11}} + \delta^4_2 \frac{\partial z^4_2}{\partial w^3_{11}} \\ \phantom{\frac{\partial c}{\partial w^3_{11}}} = \delta^4_1 w^4_{11} \sigma'(z^3_1) a^2_1 + \delta^4_2 w^4_{21} \sigma'(z^3_1) a^2_1\\ \end{array}\)
Now
\(\begin{array}{l} \frac{\partial c}{\partial w^3_{12}} = \frac{\partial c}{\partial a^4_1} \frac{\partial a^4_1}{\partial w^3_{12}} + \frac{\partial c}{\partial a^4_2} \frac{\partial a^4_2}{\partial w^3_{12}} \\ \phantom{\frac{\partial c}{\partial w^4_{12}}} = \frac{\partial c}{\partial a^4_1} \frac{\partial a^4_1}{\partial z^4_1} \frac{\partial z^4_1}{\partial w^3_{12}} + \frac{\partial c}{\partial a^4_2} \frac{\partial a^4_2}{\partial z^4_2} \frac{\partial z^4_2}{\partial w^3_{12}} \\ \phantom{\frac{\partial c}{\partial w^4_{12}}} = \delta^4_1 \frac{\partial z^4_1}{\partial w^3_{12}} + \delta^4_2 \frac{\partial z^4_2}{\partial w^3_{12}} \\ \end{array}\)
and
\(\begin{array}{l} \frac{\partial z^4_1}{\partial w^3_{12}} = \frac{\partial z^4_1}{\partial a^3_1} \frac{\partial a^3_1}{\partial z^3_1} \frac{\partial z^3_1}{\partial w^3_{12}} + \frac{\partial z^4_1}{\partial a^3_2} \frac{\partial a^3_2}{\partial z^3_2} \frac{\partial z^3_2}{\partial w^3_{12}}\\ \phantom{\frac{\partial z^4_1}{\partial w^3_{12}}} = w^4_{11}{\times} \sigma'(z^3_1){\times} a^2_2 + w^4_{12}{\times} \sigma'(z^3_2){\times} 0\\ \phantom{\frac{\partial z^4_1}{\partial w^3_{12}}} = w^4_{11}{\times} \sigma'(z^3_1){\times} a^2_2\\ \end{array}\)
\(\begin{array}{l} \frac{\partial z^4_2}{\partial w^3_{12}} = \frac{\partial z^4_2}{\partial a^3_1} \frac{\partial a^3_1}{\partial z^3_1} \frac{\partial z^3_1}{\partial w^3_{12}} + \frac{\partial z^4_2}{\partial a^3_2} \frac{\partial a^3_2}{\partial z^3_2} \frac{\partial z^3_2}{\partial w^3_{12}}\\ \phantom{\frac{\partial z^4_2}{\partial w^3_{12}}} = w^4_{21}{\times} \sigma'(z^3_1){\times} a^2_2 + w^4_{22}{\times} \sigma'(z^3_2){\times} 0\\ \phantom{\frac{\partial z^4_2}{\partial w^3_{12}}} = w^4_{21}{\times} \sigma'(z^3_1){\times} a^2_2\\ \end{array}\)
Hence:
\(\begin{array}{l} \frac{\partial c}{\partial w^3_{12}} = \delta^4_1 \frac{\partial z^4_1}{\partial w^3_{12}} + \delta^4_2 \frac{\partial z^4_2}{\partial w^3_{12}} \\ \phantom{\frac{\partial c}{\partial w^3_{12}}} = \delta^4_1 w^4_{11} \sigma'(z^3_1) a^2_2 + \delta^4_2 w^4_{21} \sigma'(z^3_1) a^2_2\\ \end{array}\)
Now the derivatives with respect to the bottom neurons in layer 3 are
\(\begin{array}{l} \frac{\partial c}{\partial w^3_{21}} = \frac{\partial c}{\partial a^4_1} \frac{\partial a^4_1}{\partial w^3_{21}} + \frac{\partial c}{\partial a^4_2} \frac{\partial a^4_2}{\partial w^3_{21}} \\ \phantom{\frac{\partial c}{\partial w^4_{11}}} = \frac{\partial c}{\partial a^4_1} \frac{\partial a^4_1}{\partial z^4_1} \frac{\partial z^4_1}{\partial w^3_{21}} + \frac{\partial c}{\partial a^4_2} \frac{\partial a^4_2}{\partial z^4_2} \frac{\partial z^4_2}{\partial w^3_{21}} \\ \phantom{\frac{\partial c}{\partial w^4_{11}}} = \delta^4_1 \frac{\partial z^4_1}{\partial w^3_{21}} + \delta^4_2 \frac{\partial z^4_2}{\partial w^3_{21}} \\ \end{array}\)
and
\(\begin{array}{l} \frac{\partial z^4_1}{\partial w^3_{21}} = \frac{\partial z^4_1}{\partial a^3_1} \frac{\partial a^3_1}{\partial z^3_1} \frac{\partial z^3_1}{\partial w^3_{21}} + \frac{\partial z^4_1}{\partial a^3_2} \frac{\partial a^3_2}{\partial z^3_2} \frac{\partial z^3_2}{\partial w^3_{21}}\\ \phantom{\frac{\partial z^4_1}{\partial w^3_{21}}} = w^4_{11}{\times} \sigma'(z^3_1){\times} 0 + w^4_{12}{\times} \sigma'(z^3_2){\times} a^2_1\\ \phantom{\frac{\partial z^4_1}{\partial w^3_{21}}} = w^4_{12}{\times} \sigma'(z^3_2){\times} a^2_1\\ \end{array}\)
\(\begin{array}{l} \frac{\partial z^4_2}{\partial w^3_{21}} = \frac{\partial z^4_2}{\partial a^3_1} \frac{\partial a^3_1}{\partial z^3_1} \frac{\partial z^3_1}{\partial w^3_{21}} + \frac{\partial z^4_2}{\partial a^3_2} \frac{\partial a^3_2}{\partial z^3_2} \frac{\partial z^3_2}{\partial w^3_{21}}\\ \phantom{\frac{\partial z^4_1}{\partial w^3_{21}}} = w^4_{21}{\times} \sigma'(z^3_1){\times} 0 + w^4_{22}{\times} \sigma'(z^3_2){\times} a^2_1\\ \phantom{\frac{\partial z^4_1}{\partial w^3_{21}}} = w^4_{22}{\times} \sigma'(z^3_2){\times} a^2_1\\ \end{array}\)
Hence:
\(\begin{array}{l} \frac{\partial c}{\partial w^3_{21}} = \delta^4_1 \frac{\partial z^4_1}{\partial w^3_{21}} + \delta^4_2 \frac{\partial z^4_2}{\partial w^3_{21}} \\ \phantom{\frac{\partial c}{\partial w^4_{11}}} = \delta^4_1 w^4_{12} \sigma'(z^3_2) a^2_1 + \delta^4_2 w^4_{22} \sigma'(z^3_2) a^2_1\\ \end{array}\)
Now
\(\begin{array}{l} \frac{\partial c}{\partial w^3_{22}} = \frac{\partial c}{\partial a^4_1} \frac{\partial a^4_1}{\partial w^3_{22}} + \frac{\partial c}{\partial a^4_2} \frac{\partial a^4_2}{\partial w^3_{22}} \\ \phantom{\frac{\partial c}{\partial w^4_{11}}} = \frac{\partial c}{\partial a^4_1} \frac{\partial a^4_1}{\partial z^4_1} \frac{\partial z^4_1}{\partial w^3_{22}} + \frac{\partial c}{\partial a^4_2} \frac{\partial a^4_2}{\partial z^4_2} \frac{\partial z^4_2}{\partial w^3_{22}} \\ \phantom{\frac{\partial c}{\partial w^4_{11}}} = \delta^4_1 \frac{\partial z^4_1}{\partial w^3_{22}} + \delta^4_2 \frac{\partial z^4_2}{\partial w^3_{22}}\\ \end{array}\)
and
\(\begin{array}{l} \frac{\partial z^4_1}{\partial w^3_{22}} = \frac{\partial z^4_1}{\partial a^3_1} \frac{\partial a^3_1}{\partial z^3_1} \frac{\partial z^3_1}{\partial w^3_{22}} + \frac{\partial z^4_1}{\partial a^3_2} \frac{\partial a^3_2}{\partial z^3_2} \frac{\partial z^3_2}{\partial w^3_{22}}\\ \phantom{\frac{\partial z^4_1}{\partial w^3_{22}}} = w^4_{11}{\times} \sigma'(z^3_1){\times} 0 + w^4_{12}{\times} \sigma'(z^3_2){\times} a^2_2\\ \phantom{\frac{\partial z^4_1}{\partial w^3_{22}}} = w^4_{12}{\times} \sigma'(z^3_2){\times} a^2_2\\ \end{array}\)
\(\begin{array}{l} \frac{\partial z^4_2}{\partial w^3_{22}} = \frac{\partial z^4_2}{\partial a^3_1} \frac{\partial a^3_1}{\partial z^3_1} \frac{\partial z^3_1}{\partial w^3_{22}} + \frac{\partial z^4_2}{\partial a^3_2} \frac{\partial a^3_2}{\partial z^3_2} \frac{\partial z^3_2}{\partial w^3_{22}}\\ \phantom{\frac{\partial z^4_2}{\partial w^3_{22}}} = w^4_{21}{\times} \sigma'(z^3_1){\times} 0 + w^4_{22}{\times} \sigma'(z^3_2){\times} a^2_2\\ \phantom{\frac{\partial z^4_2}{\partial w^3_{22}}} = w^4_{22}{\times} \sigma'(z^3_2){\times} a^2_2\\ \end{array}\)
Hence:
\(\begin{array}{l} \frac{\partial c}{\partial w^3_{22}} = \frac{\partial c}{\partial a^4_1} \frac{\partial a^4_1}{\partial w^3_{22}} + \frac{\partial c}{\partial a^4_2} \frac{\partial a^4_2}{\partial w^3_{22}} \\ \phantom{\frac{\partial c}{\partial w^4_{11}}} = \delta^4_1 w^4_{12} \sigma'(z^3_2) a^2_2 + \delta^4_2 w^4_{22} \sigma'(z^3_2) a^2_2\\ \end{array}\)
Rearranging the equations for \(\frac{\partial c}{\partial w^3_{jk}}\) derived above:
\(\begin{array}{ll} \begin{array}{l} \frac{\partial c}{\partial w^3_{11}} = (\delta^4_1 w^4_{11} + \delta^4_2 w^4_{21}) \sigma'(z^3_1) a^2_1\\ \end{array} & \begin{array}{l} \frac{\partial c}{\partial w^3_{12}} = (\delta^4_1 w^4_{11} + \delta^4_2 w^4_{21}) \sigma'(z^3_1) a^2_2\\ \end{array} \\ \begin{array}{l} \frac{\partial c}{\partial w^3_{21}} = (\delta^4_1 w^4_{12} + \delta^4_2 w^4_{22}) \sigma'(z^3_2) a^2_1\\ \end{array} & \begin{array}{l} \frac{\partial c}{\partial w^3_{22}} = (\delta^4_1 w^4_{12} + \delta^4_2 w^4_{22}) \sigma'(z^3_2) a^2_2\\ \end{array} \end{array}\)
Using
\(\begin{array}{l} \delta^3_1 = (\delta^4_1 w^4_{11} + \delta^4_2 w^4_{21}) \sigma'(z^3_1)\\ \end{array}\) \(\qquad\) \(\begin{array}{l} \delta^3_2 = (\delta^4_1 w^4_{12} + \delta^4_2 w^4_{22}) \sigma'(z^3_2)\\ \end{array}\)
the equations for \(\frac{\partial c}{\partial w^3_{jk}}\) can be shortened to:
\(\begin{array}{ll} \begin{array}{l} \frac{\partial c}{\partial w^3_{11}} = \delta^3_1 a^2_1\\ \end{array} & \begin{array}{l} \frac{\partial c}{\partial w^3_{12}} = \delta^3_1 a^2_2\\ \end{array} \\ \begin{array}{l} \frac{\partial c}{\partial w^3_{21}} = \delta^3_2 a^2_1\\ \end{array} & \begin{array}{l} \frac{\partial c}{\partial w^3_{22}} = \delta^3_2 a^2_2\\ \end{array} \end{array}\)
Now calculate \(\frac{\partial c}{\partial b^3_j}\)
\(\begin{array}{l} \frac{\partial c}{\partial b^3_j} = \frac{\partial c}{\partial a^4_1} \frac{\partial a^4_1}{\partial b^3_j} + \frac{\partial c}{\partial a^4_2} \frac{\partial a^4_2}{\partial b^3_j}\\ \phantom{\frac{\partial c}{\partial b^3_j}} = \frac{\partial c}{\partial a^4_1} \frac{\partial a^4_1}{\partial z^4_1} \frac{\partial z^4_1}{\partial b^3_j} + \frac{\partial c}{\partial a^4_2} \frac{\partial a^4_2}{\partial z^4_2} \frac{\partial z^4_2}{\partial b^3_j}\\ \phantom{\frac{\partial c}{\partial b^3_j}} = \delta^4_1 \frac{\partial z^4_1}{\partial b^3_j} + \delta^4_2 \frac{\partial z^4_2}{\partial b^3_j}\\ \end{array}\)
and
\(\begin{array}{l} \frac{\partial z^4_1}{\partial b^3_j} = \frac{\partial z^4_1}{\partial a^3_1} \frac{\partial a^3_1}{\partial z^3_1} \frac{\partial z^3_1}{\partial b^3_j} + \frac{\partial z^4_1}{\partial a^3_2} \frac{\partial a^3_2}{\partial z^3_2} \frac{\partial z^3_2}{\partial b^3_j} = w^4_{11} \sigma'(z^3_1) \frac{\partial z^3_1}{\partial b^3_j} + w^4_{12} \sigma'(z^3_2) \frac{\partial z^3_2}{\partial b^3_j} \end{array}\)
\(\begin{array}{l} \frac{\partial z^4_2}{\partial b^3_j} = \frac{\partial z^4_2}{\partial a^3_1} \frac{\partial a^3_1}{\partial z^3_1} \frac{\partial z^3_1}{\partial b^3_j} + \frac{\partial z^4_2}{\partial a^3_2} \frac{\partial a^3_2}{\partial z^3_2} \frac{\partial z^3_2}{\partial b^3_j} = w^4_{21} \sigma'(z^3_1) \frac{\partial z^3_1}{\partial b^3_j} + w^4_{22} \sigma'(z^3_2) \frac{\partial z^3_2}{\partial b^3_j} \end{array}\)
So:
\(\begin{array}{l} \frac{\partial c}{\partial b^3_j} = \delta^4_1 (w^4_{11} \sigma'(z^3_1) \frac{\partial z^3_1}{\partial b^3_j} + w^4_{12} \sigma'(z^3_2) \frac{\partial z^3_2}{\partial b^3_j}) + \delta^4_2 (w^4_{21} \sigma'(z^3_1) \frac{\partial z^3_1}{\partial b^3_j} + w^4_{22} \sigma'(z^3_2) \frac{\partial z^3_2}{\partial b^3_j}) \end{array}\)
Taking \(j=1\) and \(j=2\) and doing obvious simplifications:
\(\begin{array}{l} \frac{\partial c}{\partial b^3_1} = \delta^4_1 (w^4_{11} \sigma'(z^3_1) \frac{\partial z^3_1}{\partial b^3_1} + w^4_{12} \sigma'(z^3_2) \frac{\partial z^3_2}{\partial b^3_1}) + \delta^4_2 (w^4_{21} \sigma'(z^3_1) \frac{\partial z^3_1}{\partial b^3_1} + w^4_{22} \sigma'(z^3_2) \frac{\partial z^3_2}{\partial b^3_1})\\ \phantom{\frac{\partial c}{\partial b^3_1}} = \delta^4_1 (w^4_{11} \sigma'(z^3_1) 1 + w^4_{12} \sigma'(z^3_2) 0) + \delta^4_2 (w^4_{21} \sigma'(z^3_1) 1 + w^4_{22} \sigma'(z^3_2) 0)\\ \phantom{\frac{\partial c}{\partial b^3_1}} = \delta^4_1 w^4_{11} \sigma'(z^3_1) + \delta^4_2 w^4_{21} \sigma'(z^3_1)\\ \phantom{\frac{\partial c}{\partial b^3_1}} = (\delta^4_1 w^4_{11} + \delta^4_2 w^4_{21}) \sigma'(z^3_1)\\ \phantom{\frac{\partial c}{\partial b^3_1}} = \delta^3_1\\ \end{array}\)
\(\begin{array}{l} \frac{\partial c}{\partial b^3_2} = \delta^4_1 (w^4_{11} \sigma'(z^3_1) \frac{\partial z^3_1}{\partial b^3_2} + w^4_{12} \sigma'(z^3_2) \frac{\partial z^3_2}{\partial b^3_2}) + \delta^4_2 (w^4_{21} \sigma'(z^3_1) \frac{\partial z^3_1}{\partial b^3_2} + w^4_{22} \sigma'(z^3_2) \frac{\partial z^3_2}{\partial b^3_2})\\ \phantom{\frac{\partial c}{\partial b^3_2}} = \delta^4_1 (w^4_{11} \sigma'(z^3_1) 0 + w^4_{12} \sigma'(z^3_2) 1) + \delta^4_2 (w^4_{21} \sigma'(z^3_1) 0 + w^4_{22} \sigma'(z^3_2) 1)\\ \phantom{\frac{\partial c}{\partial b^3_2}} = \delta^4_1 w^4_{12} \sigma'(z^3_2) + \delta^4_2 w^4_{22} \sigma'(z^3_2)\\ \phantom{\frac{\partial c}{\partial b^3_2}} = (\delta^4_1 w^4_{12} + \delta^4_2 w^4_{22}) \sigma'(z^3_2)\\ \phantom{\frac{\partial c}{\partial b^3_2}} = \delta^3_2\\ \end{array}\)
The equations for \(\delta^l_j\), \(\frac{\partial c}{\partial w^l_{jk}}\) and \(\frac{\partial c}{\partial b^l_j}\) with \(l=4\) and \(l=3\) are derived above.
The equations for \(l=2\) and \(l=1\) are derived in a completely analogous manner: just replay the calculations with smaller superscripts. The complete set of equations are given in the table below.
\(\begin{array}{|l|l|l|} \hline \begin{array}{l} \delta^4_1 = (a^4_1 - y_1) \sigma'(z^4_1) \end{array} & \begin{array}{l} \delta^4_2 = (a^4_2 - y_2) \sigma'(z^4_2)\\ \end{array} \\ \hline \begin{array}{l} \frac{\partial c}{\partial w^4_{11}} = \delta^4_1 a^3_1 \end{array} & \begin{array}{l} \frac{\partial c}{\partial w^4_{21}} = \delta^4_2 a^3_1 \\ \end{array} & \begin{array}{l} \frac{\partial c}{\partial b^4_1} = \delta^4_1 \end{array} \\ \hline \begin{array}{l} \frac{\partial c}{\partial w^4_{12}} = \delta^4_1 a^3_2 \end{array} & \begin{array}{l} \frac{\partial c}{\partial w^4_{22}} = \delta^4_2 a^3_2 \\ \end{array} & \begin{array}{l} \frac{\partial c}{\partial b^4_2} = \delta^4_2\\ \end{array} \\ \hline \hline \begin{array}{l} \delta^3_1 = (\delta^4_1 w^4_{11} + \delta^4_2 w^4_{21}) \sigma'(z^3_1) \end{array} & \begin{array}{l} \delta^3_2 = (\delta^4_1 w^4_{12} + \delta^4_2 w^4_{22}) \sigma'(z^3_2)\\ \end{array} \\ \hline \begin{array}{l} \frac{\partial c}{\partial w^3_{11}} = \delta^3_1 a^2_1\\ \end{array} & \begin{array}{l} \frac{\partial c}{\partial w^3_{12}} = \delta^3_1 a^2_2\\ \end{array} & \begin{array}{l} \frac{\partial c}{\partial b^3_1} = \delta^3_1\\ \end{array} \\ \hline \begin{array}{l} \frac{\partial c}{\partial w^3_{21}} = \delta^3_2 a^2_1\\ \end{array} & \begin{array}{l} \frac{\partial c}{\partial w^3_{22}} = \delta^3_2 a^2_2\\ \end{array} & \begin{array}{l} \frac{\partial c}{\partial b^3_2} = \delta^3_2\\ \end{array} \\ \hline \hline \begin{array}{l} \delta^2_1 = (\delta^3_1 w^3_{11} + \delta^3_2 w^3_{21}) \sigma'(z^2_1) \end{array} & \begin{array}{l} \delta^2_2 = (\delta^3_1 w^3_{12} + \delta^3_2 w^3_{22}) \sigma'(z^2_2)\\ \end{array} \\ \hline \begin{array}{l} \frac{\partial c}{\partial w^2_{11}} = \delta^2_1 a^1_1\\ \end{array} & \begin{array}{l} \frac{\partial c}{\partial w^2_{12}} = \delta^2_1 a^1_2\\ \end{array} & \begin{array}{l} \frac{\partial c}{\partial b^2_1} = \delta^2_1\\ \end{array} \\ \hline \begin{array}{l} \frac{\partial c}{\partial w^2_{21}} = \delta^2_2 a^1_1\\ \end{array} & \begin{array}{l} \frac{\partial c}{\partial w^2_{22}} = \delta^2_2 a^1_2\\ \end{array} & \begin{array}{l} \frac{\partial c}{\partial b^2_2} = \delta^2_2\\ \end{array} \\ \hline \hline \begin{array}{l} \delta^1_1 = (\delta^2_1 w^2_{11} + \delta^2_2 w^2_{21}) \sigma'(z^1_1) \end{array} & \begin{array}{l} \delta^1_2 = (\delta^2_1 w^2_{12} + \delta^2_2 w^2_{22}) \sigma'(z^1_2)\\ \end{array} \\ \hline \begin{array}{l} \frac{\partial c}{\partial w^1_{11}} = \delta^1_1 a^0_1\\ \end{array} & \begin{array}{l} \frac{\partial c}{\partial w^1_{12}} = \delta^1_1 a^0_2\\ \end{array} & \begin{array}{l} \frac{\partial c}{\partial b^1_1} = \delta^1_1\\ \end{array} \\ \hline \begin{array}{l} \frac{\partial c}{\partial w^1_{21}} = \delta^1_2 a^0_1\\ \end{array} & \begin{array}{l} \frac{\partial c}{\partial w^1_{22}} = \delta^1_2 a^0_2\\ \end{array} & \begin{array}{l} \frac{\partial c}{\partial b^1_2} = \delta^1_2\\ \end{array} \\ \hline \end{array}\)
Recall the vector equation \(a^l = \sigma(w^l \times a^{(l-1)} + b^l)\) where:
\(a^l = \left\lbrack \begin{matrix} a^l_1 \cr a^l_2 \cr \end{matrix}\right\rbrack\) \(\textrm{and}~\) \(w^l = \left\lbrack \begin{matrix} w^l_{11} & w^l_{12} \cr w^l_{21} & w^l_{22} \cr \end{matrix}\right\rbrack\) \(\textrm{and}~\) \(a^{(l-1)} = \left\lbrack \begin{matrix} a^{(l-1)}_1 \cr a^{(l-1)}_2 \cr \end{matrix}\right\rbrack\) \(\textrm{and}~\) \(b^l = \left\lbrack \begin{matrix} b^l_1 \cr b^l_2 \cr \end{matrix}\right\rbrack\).
Let \({w^l}^{\top}\) be the transpose of \(w^l\), \(z^l\) the vector with components \(z^l_j\), \(\delta^l\) the vector with components \(\delta^l_j\) and \(y\) the vector with components \(y_j\), so:
\({w^l}^{\top} = \left\lbrack \begin{matrix} w^l_{11} & w^l_{21} \cr w^l_{12} & w^l_{22} \cr \end{matrix}\right\rbrack\) \(\textrm{and}~\) \(z^l = \left\lbrack \begin{matrix} z^l_1 \cr z^l_2 \cr \end{matrix}\right\rbrack\) \(\textrm{and}~\) \(\delta^l = \left\lbrack \begin{matrix} \delta^l_1 \cr \delta^l_2 \cr \end{matrix}\right\rbrack\) \(\textrm{and}~\) \(y = \left\lbrack \begin{matrix} \vphantom{z^l_1} y_1 \cr \vphantom{z^l_2} y_2 \cr \end{matrix}\right\rbrack\).
If \(u\) ans \(v\) are vectors, then \(u \odot v\) is their elementwise product (also called the Hadamard product) and \(u+v\) and \(u-v\) their elementwise sum and difference, respectively.
\(\left\lbrack \begin{matrix} u_1 \cr u_2 \cr \vdots \cr u_r \cr \end{matrix}\right\rbrack \odot \left\lbrack \begin{matrix} v_1 \cr v_2 \cr \vdots \cr v_r \cr \end{matrix}\right\rbrack = \left\lbrack \begin{matrix} u_1{\times}v_1 \cr u_2{\times}v_2 \cr \vdots \cr u_r{\times}v_r \cr \end{matrix}\right\rbrack\) \(~\mathrm{and}~\) \(\left\lbrack \begin{matrix} u_1 \cr u_2 \cr \vdots \cr u_r \cr \end{matrix}\right\rbrack + \left\lbrack \begin{matrix} v_1 \cr v_2 \cr \vdots \cr v_r \cr \end{matrix}\right\rbrack = \left\lbrack \begin{matrix} u_1{+}v_1 \cr u_2{+}v_2 \cr \vdots \cr u_r{+}v_r \cr \end{matrix}\right\rbrack\).
Recall that \(z^l_j = (\sum\limits_k w^l_{jk} a^{l{-}1}_k) + b^l_j\) so, for example:
\(z^3_1 = (w^3_{11} a^{2}_1 + w^3_{12} a^{2}_2) + b^3_1\) \(~\mathrm{and}~\) \(z^3_2 = (w^3_{21} a^{2}_1 + w^3_{22} a^{2}_2) + b^3_2\)
These two equations can be written as the single equation \(z^3 = w^3 a^2 + b^3\) using vectors and matrices.
The equations for \(\delta^3_1\) and \(\delta^3_2\) are
\(\delta^3_1 = (\delta^4_1 w^4_{11} + \delta^4_2 w^4_{21}) \sigma'(z^3_1)\) \(~\mathrm{and}~\) \(\delta^3_2 = (\delta^4_1 w^4_{12} + \delta^4_2 w^4_{22}) \sigma'(z^3_2)\)
which – noting that as multiplication is commutative so \(\delta^l_j w^l_{jk} = w^l_{jk} \delta^l_j\) – can be written as
\(\left\lbrack \begin{matrix} \delta^3_1 \cr \delta^3_2 \cr \end{matrix}\right\rbrack = \left( \left\lbrack \begin{matrix} w^4_{11} & w^4_{21} \cr w^4_{12} & w^4_{22} \cr \end{matrix}\right\rbrack \times \left\lbrack \begin{matrix} \delta^4_1 \cr \delta^4_2 \cr \end{matrix}\right\rbrack \right) \odot \sigma'\hskip-1mm \left( \left\lbrack \begin{matrix} z^3_1 \cr z^3_2 \cr \end{matrix}\right\rbrack \right)\)
which is the single vector equation \(\delta^3 = ((w^4)^{\top} \delta^4)\odot \sigma'(z^3)\).
The notation \(\nabla_{{\hskip-1mm}a^l}~c\) denotes the vector whose components are the partial derivatives \(\frac{\partial c}{\partial a^l_j}\) and so \(\nabla_{{\hskip-1mm}a^L}~c\) is the vector of the rate of changes of \(c\) with respect to each output activation. \(\nabla_{{\hskip-1mm}a}~c\) will abbreviate \(\nabla_{{\hskip-1mm}a^L}~c\). For the example:
\(\nabla_{{\hskip-1mm}a^l}~c = \left\lbrack \begin{matrix} \frac{\partial c}{\partial a^l_1} \cr \frac{\partial c}{\partial a^l_2} \cr \end{matrix}\right\rbrack\) \(~\mathrm{and}~\) \(\nabla_{{\hskip-1mm}a}~c = \left\lbrack \begin{matrix} \frac{\partial c}{\partial a^4_1} \cr \frac{\partial c}{\partial a^4_2} \cr \end{matrix}\right\rbrack = \left\lbrack \begin{matrix} \vphantom{\frac{\partial c}{\partial a^4_1}} a^4_1{-}y_1 \cr \vphantom{\frac{\partial c}{\partial a^4_2}} a^4_2{-}y_2 \cr \end{matrix}\right\rbrack\)
Armed with these notations, the table above can be used to verify the following four equations, which are called “the equations of propagation” in Chapter 2 of Nielsen’s book and named by him BP1, BP2, BP3 and BP4.
\(\begin{array}{ll} \delta^L = \nabla_{{\hskip-1mm}a}~c \odot \sigma'(z^L) & \qquad\mathrm{(BP1)} \\ & \\ \delta^l = ((w^{l+1})^{\top} \delta^{l+1}) \odot \sigma'(z^l) & \qquad\mathrm{(BP2)} \\ & \\ \frac{\partial c}{\partial b^l_j} = \delta^l_j & \qquad\mathrm{(BP3)} \\ & \\ \frac{\partial c}{\partial w^l_{jk}} = a^{l-1}_k \delta^l_j & \qquad\mathrm{(BP4)} \\ \end{array}\)
The vector equations for the activations are:
\(a^l = \sigma(w^l \times a^{(l-1)} + b^l)\) where:
so, bearing in mind that \(a^0\) is the vector of the two inputs \(x_1\) and \(x_2\), the forward pass pseudocode for the two-neuron-per-layer example is:
\(~~~\begin{array}{ll} a^1 &=~~~ \sigma(w^1 a^0 + b^1)\texttt{;} \\ a^2 &=~~~ \sigma(w^2 a^1 + b^2)\texttt{;} \\ a^3 &=~~~ \sigma(w^2 a^2 + b^3)\texttt{;} \\ a^4 &=~~~ \sigma(w^4 a^3 + b^4)\texttt{;} \\ \end{array}\)
The pseudocode for calculating the weight and bias follows - remember that \(y\) is the vector of desired outputs, i.e. \(g(x_1,x_2)\).
\(\begin{array}{lll} \delta^4 = (a^4 - y) \odot \sigma'(z^4) \texttt{;} & {\Delta}w^4_{jk} = - \eta\hskip1mm \delta^4_j a^{3}_k \texttt{;} & {\Delta}b^4_j = - \eta\hskip1mm \delta^4_j \texttt{;} \\ \delta^3 = ((w^{4})^{\top} \delta^{4}) \odot \sigma'(z^3) \texttt{;} & {\Delta}w^3_{jk} = - \eta\hskip1mm \delta^3_j a^2_k \texttt{;} & {\Delta}b^3_j = - \eta\hskip1mm \delta^3_j \texttt{;} \\ \delta^2 = ((w^{3})^{\top} \delta^{3}) \odot \sigma'(z^2) \texttt{;} & {\Delta}w^2_{jk} = - \eta\hskip1mm \delta^2_j a^1_k \texttt{;} & {\Delta}b^2_j = - \eta\hskip1mm \delta^2_j \texttt{;} \\ \delta^1 = ((w^{2})^{\top} \delta^{2}) \odot \sigma'(z^1) \texttt{;} & {\Delta}w^1_{jk} = - \eta\hskip1mm \delta^1_j a^0_k \texttt{;} & {\Delta}b^1_j = - \eta\hskip1mm \delta^1_j \texttt{;} \\ \end{array}\)
Compare this with the scalar-calculation pseudocode for the one-neuron-per-layer example deltas calculation (remember that for this subscripts index layers, whereas in the two-neuron-per-layer example superscripts index layers).
\(\begin{array}{lll} \delta_4 = (a_4 - g~x)\sigma'(w_4 a_3 + b_4) \texttt{;} & {\Delta}w_4 = - \eta\hskip1mm \delta_4 a_3 \texttt{;} & {\Delta}b_4 = - \eta\hskip1mm \delta_4 \texttt{;} \\ \delta_3 = \delta_4 \sigma'(w_3 a_2 + b_3) w_4 \texttt{;} & {\Delta}w_3 = - \eta\hskip1mm \delta_3 a_2 \texttt{;} & {\Delta}b_3 = - \eta\hskip1mm \delta_3 \texttt{;} \\ \delta_2 = \delta_3 \sigma'(w_2 a_1 + b_2) w_3 \texttt{;} & {\Delta}w_2 = - \eta\hskip1mm \delta_2 a_1 \texttt{;} & {\Delta}b_2 = - \eta\hskip1mm \delta_2 \texttt{;} \\ \delta_1 = \delta_2 \sigma'(w_1 x + b_1) w_2 \texttt{;} & {\Delta}w_1 = - \eta\hskip1mm \delta_1 x \texttt{;} & {\Delta}b_1 = - \eta\hskip1mm \delta_1 \texttt{;} \\ \end{array}\)
Generalising from two neurons per layer to many neurons per layer is just a matter of increasing the range of the subscripts \(j\) and \(k\). Similarly increasing the number of layers is just a matter of increasing the range of \(l\). In the two-neuron-per-layer example
\(z^l_j = (\sum\limits_{k=1}^{k=2} w^l_{jk} a^{l{-}1}_k) + b^l_j\)
where \(j\in\{1,2\}\), \(l\in\{1,2,3,4\}\). If there are \(K\) neurons per layer and \(L\) layers, then the equation becomes:
\(z^l_j = (\sum\limits_{k=1}^{k=K} w^l_{jk} a^{l{-}1}_k) + b^l_j\)
and \(j\in\{1,\ldots,K\}\), \(l\in\{1,\ldots,L\}\).
If the equation is written in vector form, so the sum \(\Sigma\) is assimilated into a matrix multiplication, then only the number-of-layers superscript remains: \(z^l = w^l a^{l{-}1} + b^l\) and the number of neurons per layer is implicit.
The vectorised pseudocode in the general case is then as follows.
\(~~~\begin{array}{ll} a^1 &=~~~ \sigma(w^1 a^0 + b^1)\texttt{;} \\ \vdots & \phantom{=~~~} \vdots \\ a^l &=~~~ \sigma(w^l a^{l{-}1} + b^l)\texttt{;} \\ \vdots & \phantom{=~~~} \vdots \\ a^L &=~~~ \sigma(w^L a^{L{-}1} + b^L)\texttt{;} \\ \end{array}\)
The pseudocode for calculating the weight and bias follows - remember that \(y\) is the vector of desired outputs, i.e. \(g(x_1,\ldots,x_K)\). The algorithm iteratively decrements \(l\) from \(L\) down to \(1\):
\(\begin{array}{lll} \delta^L = (a^L - y) \odot \sigma'(w^L a^{L{-}1} + b^L) \texttt{;} & {\Delta}w^L_{jk} = - \eta\hskip1mm \delta^L_j a^{L{-}1}_k \texttt{;} & {\Delta}b^L_j = - \eta\hskip1mm \delta^L_j \texttt{;} \\ ~\vdots & ~\vdots \\ \delta^l = ((w^{l{+}1})^{\top} \delta^{l{+}1}) \odot \sigma'(w^l a^{l{-}1} + b^l) \texttt{;} & {\Delta}w^l_{jk} = - \eta\hskip1mm \delta^l_j a^{l{-}1}_k \texttt{;} & {\Delta}b^l_j = - \eta\hskip1mm \delta^l_j \texttt{;} \\ ~\vdots & ~\vdots \\ \delta^1 = ((w^{2})^{\top} \delta^{2}) \odot \sigma'(w^1 a^{0} + b^1) \texttt{;} & {\Delta}w^1_{jk} = - \eta\hskip1mm \delta^1_j a^0_k \texttt{;} & {\Delta}b^1_j = - \eta\hskip1mm \delta^1_j \texttt{;} \\ \end{array}\)
So far all the layers have had the same number of neurons. At the start of the section above entitles The Backpropagation algorithm, the example network shown in the diagram had different numbers of neurons in each layer. There are applications where different layers have different numbers of neurons. An intriguing discussion of one application that uses a small hidden layer is called “autocoding” by Winston and discussed in Lecture 12b of his online MIT AI course beginning about 16 minutes after the start of the YouTube video. The backpropagation algorithm for nets whose layers vary in size isn’t considered here, but I imagine that it’s a straightforward adaption of the ideas described above.
I wrote this with the limited goal of teaching myself how backpropagation works. The bigger picture is presented in numerous online sources, as discussed earlier, and there’s a peek at recent stuff from Google and its cool applications, ranging from cucumber classification to cryptography, in Martin Abadi’s keynote talk at ICPF 2016.
I was hoping to finish by making a few general comments on machine learning, artificial intelligence, the singularity and so on … but right now am feeling burned out on derivatives and matrices. Maybe I’ll add more thoughts when I’ve recovered.